Fulvia Gloria-Bottini
Department of Biomedicine and Prevention, University of Tor Vergata, Via Montpellier, 00133 Rome (Italy)
Phone: +39 06 30889514
E-mail address : gloria@med.uniroma2.it
© 2019 Sift Desk Journals. All Rights Reserved
VOLUME: 3 ISSUE: 1
Fulvia Gloria-Bottini
Department of Biomedicine and Prevention, University of Tor Vergata, Via Montpellier, 00133 Rome (Italy)
Phone: +39 06 30889514
E-mail address : gloria@med.uniroma2.it
Gloria-Bottini F1, Neri A2, Saccucci P1, Manca Bitti, M L2 Rapini N2, Magrini A1, Bottini E1
Department of Biomedicine and Prevention, University of Rome Tor Vergata, School of Medicine, Rome, Italy.
Department of Medical Science, Paediatric Diabetes Unit, University of Tor Vergata, Rome, Italy
Sana Irfan Khan(drsanashaham@gmail.com)
Sarah E Harris(sarah.harris@igmm.ed.ac.uk)
Gloria-Bottini.F., IMMUNE RELATED DISEASES AND THEIR RELATIONSHIP WITH THE GENETIC VARIABILITY WITHIN THE ADENOSINE DEAMINASE GENE(2018)Journal of Advances In Allergy & Immunologic Diseases 3(1)
Background: Adenosine deaminase (ADA) structural gene consists of 12 exons. A number of differences among normal sequences have been found within the coding and intronic region of the gene: the possible role of this variability on ADA function are unknown. In the present study we reexamined our data on a set of immune related diseases [Type 1 Diabetes (T1D), Rheumatoid Arthritis (RA), Endometriosis (ENDO) and Non Diabetic Coronary Artery Diseases (NDCAD)] investigating with a new approach the relevance of ADA genetic variability on susceptibility to these disorders.
Methods: Three single nucleotide polymorphisms were considered, namely Taq1 site (ADA1) on exon 1, Pst1 site on intron 2 (ADA2) and MLu NI (ADA6) on exon 6. We reanalyzed the data on 199 children with T1D, on 79 subjects with RA, on 98 women with ENDO, on 136 NDCAD and on 246 healthy blood donors. ADA genotype was determined by DNA analysis. The subjects were classified according to the number of homozygous genotypes for the more frequent alleles at the three loci.
Results: All diseases show a strong excess of subjects with the more frequent genotype in all loci as compared to controls suggesting a protective effect of heterozygosity within the ADA gene.
Conclusions: The data suggest that the combination of polymorphic sites within the ADA gene influences the susceptibility to some immune related disorders and calls for further studies on the relationship between ADA gene structure and immune functions.
The ADA gene is composed of 12 exons and is localized on 20q13.11 [1]. Differences among normal sequences have been found both within the coding and intronic regions of gene and a study on linkage disequilibrium has shown areas of weak associations and those of strong associations within pairs of internal markers [2,3].
The presence of a Taq1 site (ADA1) (nt 4050-4053) at exon 1 corresponds to a functional variation due to a substitution of asparagine for aspartic acid at codon 8. This mutation represents the basis of the biochemical polymorphism described by Spencer et al [4] with two common alleles ADA1*1 and ADA1*2.
Adenosine deaminase catalyzes the irreversible deamination of adenosine to inosine, contributing to the regulation of adenosine concentration in body fluids. Adenosine is a local hormone that influences blood flow and platelet aggregation, increases insulin sensitivity in adipocytes [5-7] and decreases insulin sensitivity in muscle fibers [8,9]. Decreased concentration of adenosine weakens T cell activation while its high concentration increase the activity of adenosine receptors strengthening T cell activation [10,11]. Adenosine is a cardio protective agent [12] and impairment of adenosine related signal transduction contributes to chronic health failure [13].
ADA acts also as an ecto-enzyme [14,15] contributing to degradation of extracellular adenosine [11] and to the signal transduction through interaction with CD26 and A1R [16].
In the present study we investigated the possible influence of allelic combination within the ADA gene on the susceptibility to a set of immune related disorders. Three single nucleotide polymorphisms ADA1, ADA2 and ADA6 were examined [17]. The position of polymorphic sites within ADA gene and the restriction enzymes are presented in table 1. ADA1 is an exonic site involved in the regulation of enzymatic activity. ADA2 is an intronic polymorphism and ADA6 is a synonymous substitution, therefore they do not change the protein sequence but could influence tissue specific expression. Moreover, ADA2 and ADA6 polymorphisms may be markers of DNA sequences responsible for functional variation in the enzymatic activity and ecto enzymatic functions.
We studied 199 children with Type 1 Diabetes (T1D), 79 subjects with Rheumatoid Arthritis (RA), 98 women with endometriosis, 136 non diabetic subjects with Coronary Artery Disease (CAD) and 246 healthy blood donors (Controls). Data on these subjects have been previously reported [18-22]. An analysis on the role of combination of ADA1,ADA2 and ADA6 loci ,however, has not been previously performed. Informed verbal consent was obtained from the patients or from their mothers to participate in the study that was approved by the Council of our Department. The collection of data was performed a few years ago before the establishment of an Ethical Committee. All procedures followed were in accordance with the ethical standards and with the Helsinki Declaration of 1964 and ita later amendments. ADA genotypes were determined by PCR. Genomic DNA was extracted from venous blood samples collected in NaEDTA using the procedure of Kunkel [23]. PCR amplification was carried out according to Hirschorn [17] as previously described [18].
The alleles corresponding to the presence and absence of restriction site were assigned as allele *1 and allele *2 respectively.
Table 2 shows the scheme by which we classified the subjects according to the combination of ADA1, ADA2 and ADA6 genotypes. Subjects carrying three common genotypes were ranked 3 while those carrying only genotypes with the rare alleles were ranked 0.
Determination of genotypes was made in groups of twenty including cases and controls to eliminate errors due to incomplete enzymatic digestion.
Statistical analyses were performed using the SPSS package [24].
Table 3 shows the distribution of the number of homozygous genotypes for the more frequent ADA1, ADA2 and ADA6 alleles in controls and in the immune related diseases considered. All diseases show a statistically significant difference as compared to controls with a strong tendency towards an excess of homozygotes for the more frequent allele (rank 3). A border line difference (P=0.046) is observed among the diseases studied.
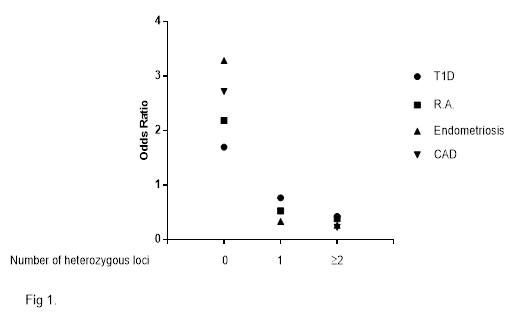
Figure 1 shows the Odds Ratio for the diseases considered in relation to the number of homozygous genotypes for the more frequent allele at the three ADA loci studied. The odds ratio for all diseases is very high in subjects who were homozygous for the more common allele in all ADA loci and decreases with the number of heterozygous loci.
The parameter proposed in the present study is an indicator of the number of mutation within the ADA gene and of the degree of heterozygosity. This new analysis is explorative: however, the results suggest that further studies in this area could be rewarding.
T1D and RA are autoimmune diseases. Immunological factors may have an important role in endometriosis [25-26] and in CAD [27-28]; furthermore the role of adenosine deaminase in immune function is well known.
We have previously performed association studies of ADA gene in T1D, in RA and in CAD. In T1D haplotype differences were observed between T1D children and controls [19]. In CAD differences between patients and controls were also observed [21,22]. Only a slight border line association was observed in RA [18].
The present approach is different from that of our previous studies and seems more productive suggesting that the combination of polymorphic sites within ADA gene influences the susceptibility to immune related diseases. The loci investigated could have a role on ADA activity and in turn on adenosine concentration. These loci could also influence the function of ADA as ecto-enzyme, thereby modulating the interaction of ADA with adenosine receptors on the surface of T cells. The association, however, could be due to other loci in linkage disequilibrium with the loci studied.
The result suggests that the heterozygosity within the ADA gene is protective against immune related diseases COMPETING INTEREST : there is any financial competing interests;
|
Name |
Position within the gene |
Restriction Enzyme |
|
ADA1 |
Exon 6 nt 4050-4053 |
Taq1 |
|
ADA2 |
Intron 2 nt 19465-19470 |
Pst1 |
|
ADA6 |
Exon 6 nt 31230-31235 |
MluN1 |
|
ADA1 |
ADA2 |
ADA6 |
Classification |
|
C |
C |
C |
3 |
|
C |
C |
R |
2 |
|
R |
C |
C |
|
|
C |
R |
C |
|
|
C |
R |
R |
1 |
|
R |
C |
R |
|
|
R |
R |
C |
|
|
R |
R |
R |
0 |
C= Homozygous for the common allele; R=Homozygous or heterozygous for the rare allele
|
|
Controls (A) |
T1D (B) |
Rheumatoid Arthritis (C) |
Endometriosis (D) |
CAD non diabetic (E) |
|
3 common genotypes |
20.7% |
30.8% |
36.4% |
46.2% |
41.5% |
|
2 common genotypes |
36.2% |
41.5% |
33.8% |
27.7% |
38.6% |
|
1 common genotype |
37.0% |
22.1% |
28.6% |
26.2% |
17.0% |
|
No common genotypes |
6.1% |
5.5% |
1.3% |
0.0% |
3.0% |
|
Total n° |
246 |
195 |
77 |
65 |
135 |
|
Chi square test of independence χ2 df p
A vs B 13.324 3 0.004 A vs C 10.413 3 0.015 A vs D 19.847 3 0.0002 A vs E 28.010 3 0.00003 B vs C vs D vs E 17.190 9 0.046
|
|||||
Viginton DA, Kaplan DJ, States JC, Akeson AL, Perme CM, Bilyk IJ, Vaughn A., Lattier DL, Hutton JJ. Complete sequenze and structure of the gene for human adenosine deaminase. Biochem. 1986; 25: 8234-8244.
View ArticleTzall S, Ellenbogen A, Eng F, Hirschhorn R. Identintification and characterization of nine RFLPs at the adenosine deaminase (ADA) locus. Am J Hum Genet. 1989; 44: 864-875. PMid:2567118
PubMed/NCBICruciani F, Bernardini L, Santolamazza P, Modiano D, Torroni A, Scozzari R. Linkage disequilibrium analysis of the human adenosine deaminase (ADA) gene provides evidence for a lack of correlation between hot spots and unequal homologous recombination. Genomics 2003; 82: 20-33. 00096-X
View ArticleSpencer N, Hopkinson D, Harris H. Adenosine deaminase polymorphism in man. Ann Hum Genet. 1968; 32: 9-14.
View ArticleLondos C, Honnor RC, Dhillon GS. Camp dependent protein kinase and lipolysis in rat adipocytes (3). Multiple modes of insulin regulation of lipolysis and regulation of insulin responses by adenylate cyclase regulators. J Biol Chem. 1985; 260:15139-15145. PMid:3905791
PubMed/NCBIOhisalo J, Strandberg H, Kostiainen E, Kuusi T, Ehnnholm C. Stimulation of lipoprotein lipase activity of rat adipose tissue and post-heparin plasma by N6- (phenylisopropyl)adenosine. FEBS Lett. 1981; 132: 121-123. 80442-5
View ArticleVannucci SJ, Nishimura H, Satoh S, Cusham SW, Holman GD, Simpson IA. Cell surface accessibility of GLUT4 glucose transporters in insulin-stimulated rat adipose cells. Modulation by isoprenaline and adenosine. Biochem. J. 1992; 288: 325-330. PMid:1445278
View Article PubMed/NCBIChallis RA, Budohoski L, McManus B, Newsholme EA. Effects of adenosine receptors antagonist on insulin resistance in soleus muscle from obese Zucker rats. Biochem. J. 1984; 21: 915-917.
View ArticleChallis RA, Richards SJ, Budohoski L. Characterization of adenosine receptor modulating insulin action in skeletal muscle. Eur J Pharmacol. 1992; 226: 121-128. 90172-R
View ArticleKameoka J, Tanaka T, Nojima Y, Schlossman SF, Morimoto C. Direct association of adenosine deaminase with a T cell activation antigen, CD26.Science. 1993; 23;261:466-9.
View ArticleDong RP, Kameoka J, Hegen M, Tanaka T, Xu Y, Schlossman SF, Morimoto C. Characterization of adenosine deaminase binding to human CD26 on T cells and its biologic role in immune response. J Immunol. 1996;156:1349-55 PMid:8568233
PubMed/NCBIBarankiewicz J, Danks AM, Abushanab E, Makings L, Wiemann T, Wallis RA, Pragnacharyulu PV, Fox A, Marangos PJ. Regulation of adenosine concentration and cytoprotective effects of novel reversible adenosine deaminase inhibitors. J Pharmacol Exp Ther. 1997; 283:1230-1238. PMid:9399998
PubMed/NCBIAsakura M, Asanuma H, Kim J, Liao Y, Nakamaru K, Fujita M, Komamura K, Isomura T, Furukawa H, Tomoike H, Kitakaze M. Impact of adenosine receptor signaling and metabolism on pathophysiology in patients with chronic heart failure. Hypertens Res. 2007; 30:781-787. PMid:18037770
View Article PubMed/NCBIFranco R, Aran JM, Colomer D, Matutes E., Vives Corrons JL. Association of adenosine deaminase with erythrocyte and platelet plasma membrane: an immunological study using light and electron microscopy. J Histochem Cytochem. 1990; 38: 653-658. PMid:2332624
View Article PubMed/NCBIFranco R, Csado V, Ciruela F, Saura C, Mallo J, Canela EI, Lluis C. Cell surface adenosine deaminase: much more than ectoenzyme. Prog Neurobiol. 1997; 52: 283-294. 00013-0
View ArticleCiruela F, Saura C, Canela EI, Mallo J, Lluis C, Franco R. Adenosine deaminase affects ligand-induced signaling by interacting with cell surface adenosine receptors. FEBS Lett 1996; 380: 219-223. 00023-3
View ArticleHirshhorn R, Yang DR, Israni A. An Asp8Asn substitution results in the adenosine deaminase (ADA) genetic polymorphism (ADA 2 allozyme): occurrence on different backgrounds and apparent intragenic crossover. Ann Hum Genet. 1994; 58: 1-9.
View ArticleSebastiani GD, Bottini N, Greco E, Saccucci P, Canu G, Lucarelli P, Gloria-Bottini F, Fontana L. A study of Adenosine-Deaminase genetic polymorphism in rheumatoid arthritis.Int J Immunopathol Pharmacol. 2010 ;23:791-5. PMid:20943049
View Article PubMed/NCBISaccucci P, Meloni GF, Verrotti A, Borgiani P, D'Annibale F, Giannini C, Lucarelli P, Bottini N, Chiarelli F, Bottini E, Gloria-Bottini F. A study of three polymorphic sites of the ADA gene in children with type 1 diabetes mellitus.J Pediatr Endocrinol Metab. 2010 ;23:283-90. PMid:20480728
View Article PubMed/NCBIGloria-Bottini F, Ammendola M, Saccucci P, Pietropolli A, Magrini A, Bottini E. The association of PTPN22 polymorphism with endometriosis: effect of genetic and clinical factors.Eur J Obstet Gynecol Reprod Biol. 2013 ;169:60-3. doi: 10.1016/j.ejogrb.2013.01.014.
View ArticleSaccucci P, Binczak-Kuleta A, Banci M, Krzysztalowska M, Dofcaci A, Safranow K, Magrini A, Loniewska B, Kornacewicz-Jach Z, Chlubek D, Bottini E, Ciechanowicz A, Gloria-Bottini F. Coronary artery disease. A study of three polymorphic sites of adenosine deaminase gene.Acta Cardiol. 2014 ;69:39-44. PMid:24640520
View Article PubMed/NCBIGloria-Bottini F, Saccucci P, Banci M, Nardi P, Scognamiglio M, Pellegrino A, Bottini E, Chiariello L. Effect of genetic factors on the association between coronary artery disease and PTPN22 polymorphism.World J Cardiol. 2014 ;6):376-80. doi: 10.4330/wjc.v6.i6.376.
View ArticleKunkel LM, Smith KD, Boyer SH, Borgaonkar DS, Wachtel SS, Miller OJ, Breg WR, Jones HW, Rary JM. Analysis of human Y-chromosome ?specific reiterated DNA in chromosome variants. Proc Natl Acad Sci. USA 1977; 74: 1245-1249.
View ArticleSPSS/PC Version 5.0. SPSS Inc., Chicago 1992.
Nothnick WB. Treating endometriosis as an autoimmune disease.Fertil Steril. 2001; 76:223-31. 01878-7
View ArticleMatarese G, De Placido G, Nikas Y, Alviggi C. Pathogenesis of endometriosis: natural immunity dysfunction or autoimmune disease? Trends Mol Med. 2003;9:223-8. 00051-0
View ArticleOrozco G, S?nchez E, Gonz?lez-Gay MA, L?pez-Nevot MA, Torres B, C?liz R, Ortego-Centeno N, Jim?nez-Alonso J, Pascual-Salcedo D, Balsa A, de Pablo R, Nu-ez-Roldan A, Gonz?lez-Escribano MF, Mart?n J. Association of a functional single-nucleotide polymorphism of PTPN22, encoding lymphoid protein phosphatase, with rheumatoid arthritis and systemic lupus erythematosus. Arthritis Rheum. 2005 ;52:219-24. PMid:15641066
View Article PubMed/NCBISaccucci P, Banci M, Cozzoli E, Neri A, Magrini A, Bottini E, Gloria-Bottini F. Atherosclerosis and PTPN22: a study in coronary artery disease. Cardiology. 2011;119:54-6. doi: 10.1159/000329919.
View Article