Raju Dash
Molecular Modeling and Drug Design Laboratory, Pharmacology Research Division, Bangladesh Council of Scientific & Industrial Research (BCSIR) Chittagong-4220, Bangladesh.
Phone: +8801815913253, Email: rajudash.bgctub@gmail.com
© 2019 Sift Desk Journals. All Rights Reserved
VOLUME: 2 ISSUE: 2
Page No: 162-171
Raju Dash
Molecular Modeling and Drug Design Laboratory, Pharmacology Research Division, Bangladesh Council of Scientific & Industrial Research (BCSIR) Chittagong-4220, Bangladesh.
Phone: +8801815913253, Email: rajudash.bgctub@gmail.com
Md. Arifuzzaman1, Sarmistha Mitra2, Tumpa Acharjee2, Tanvir Iqram Siddique2, Nurul Absar3, Raju Dash3*
aDepartment of Biotechnology and Genetic Engineering, Islamic University, Kushtia-7003, Bangladesh
bDepartment of Pharmacy, University of Chittagong, Chittagong-4331, Bangladesh
cDepartment of Biochemistry and Biotechnology, University of Science & Technology Chittagong, Chittagong- 4202, Bangladesh.
Raju Dash, Molecular Docking and Binding Free Energy Analysis of Rutin and Apigetrin as Galectin-1 Inhibitors(2018)SDRP Journal of Computational Chemistry & Molecular Modelling 2(2)
Human galectin protein 1 is one of the major carbohydrate-binding proteins and is the first member of this family which is responsible for nearly all types of cancer. Several studies have been conducted to inhibit this protein to treat cancer. However, due to the side effects of those inhibitors and also the pharmacodynamics and pharmacokinetics difficulties made these inhibitors to be less important and mostly avoided for treatment purposes. For this reason, this study was convinced to unveil the binding mechanism of two natural carbohydrate ring containing compounds, rutin and apigetrin, using computational approaches including molecular docking, binding energy calculation and ADME analysis. . According to the results, it was found that the residues, Val31, Ser29, Arg48, His44, Asn33, Glu71, Asn61 and Asp123 were involved in the interactions with the CRD domain of galectin-1. Apigetrin showed higher binding free energy than rutin in MM-GBSA binding energy calculation. ADME analysis revealed that, apigetrin showed better result than rutin for maximum parameters of ADME analysis. Taken together, apigetrin can be subjected for further analysis for the development of new inhibitor against human galectin-1 protein by in vitro and in vivo experiments.
Carbohydrate-binding protein family includes galectin-1 with a conserved carbohydrate recognition domain (CRD) responsible for β-galactoside binding with around 130 amino acids [1-3]. The galectin-1 protein is encoded by the LSGALS1 gene and is located on the 22q12 chromosome [4]. It is a monomer of 14 kda weight or a non-covalent homodimer with one CRD per subunit [4]. Presence of multiple CRDs in homodimer makes it enable to perform cell adhesion function, intracellular signaling and also forming multivalent lattices with cell surface glycoconjugates [5]. The structure and folding of galectin-1 protein contains a β sandwich with two anti-parallel β-sheets and exists as a dimer in solution [6]. Galectin-1 is responsible for cell adhesion and migration [7] and also modulates several biological and cellular processes including cell proliferation [8], apoptosis [9] and m-RNA splicing [10]. Galectin-1 was evident to be played functional roles in pathological processes such as pre-eclampsia, inflammation, diabetes, atherosclerosis and cancer [11-14].
To define tumor metastasis, altered cell adhesion, increased invasiveness and angiogenesis as well as evasion of the immune responses are well characterized [15]. Over expression of galectin-1 protein has been reported in colon cancer [16], breast cancer [17], lung cancer [18], head and neck cancer [19], ovarian cancer [20], prostate cancer [21], gliomas [22], Kaposi’s sarcoma [23], myeloproliferative neoplasia [24] and Hodgkin’s lymphoma [25]. By influencing and enhancing proangiogenic signaling pathways such as VEGF signaling, galectin-1 also regulates tumor angiogenesis [11, 26-28]. Inhibition of galectin-1 is now becomes a major target to treat cancer and several compounds were evaluated that blocks galectin-1 and its binding patterns [23, 29-34].
The major problems with these inhibitors involve the side effects of the inhibitors, for instances, synthetic lactulose amines, oligosaccharides derivatives, thiodigalactosides [29, 30, 33, 35]. Some inhibitors have high molecular weight and some with unknown affinity such as Davanat, TDG ester derivatives and anti-galectin monoclonal antibody [23, 31, 34]. The pharmacokinetics and pharmacodynamics difficulties make these inhibitors to be avoided [36]. Natural compounds are now extensively used to treat various types of diseases, and 60-70% of drugs nowadays are derived from natural sources [37]. Recently, about 30 natural compounds are in clinical trial to treat cancer [38]. There are enormous pharmacological targets and natural compounds possesses pleiotropic nature by interacting with multitargets and therefore computer aided evaluation of natural compounds are promising in recent days [39]. Lack of toxicity, low molecular weight, bioavailability and easy availability makes the natural compounds as a potential therapeutic agents [40].
By considering this issue, the present study therefore aimed to analyze new compounds rutin and apigetrin as galectin-1 inhibitors (Figure 1). Rutin, also known as rutinoside (quercetin-3-O-rutinoside) is a natural flavonol glycoside consists of quercetin and disaccharide rutinose (α-L-rhamnopyranosyl-(1-6)-β-D-glucopyranose) have multitargeted mechanism to treat cancer [41]. Rutin reported to exhibit several biochemical and pharmacological activities like free radical scavenging activity [42], reduction of inflammatory responses and anticarcinogenic properties [43]. Apigetrin, or apigetrin-7-O-glucoside, a monosaccharide derivative, has already been reported to inhibit cancer in previous study by analyzing binding free energy calculation and molecular docking against several drug targets [44]. Although monosaccharides have lower affinity than di or oligosaccharides to galectins [45, 46], however, monosaccharides based inhibitors provide higher ligand efficiency and glycolytic property and could increase bioavailability and uptake [47]. As results, this study considered both di and monosaccharids to evaluate their affinity and ligand binding efficiency as well as their inhibitory activity against galectin 1 protein by molecular docking analysis, binding free energy calculation by MM-GBSA analysis and pharmacokinetics properties analysis by ADME.
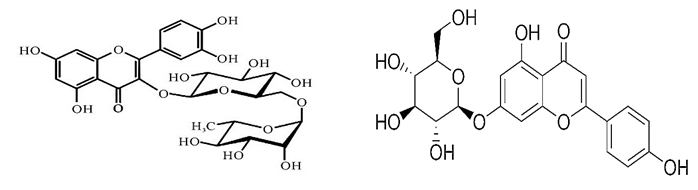
Figure 1: Two dimensional structure of rutin (Left) and apigetrin (Right)
2.1 Protein Preparation
In order to perform molecular docking analysis, at first, retrieval of the three dimensional crystal structure of the human galectin-1 (PDB ID 3T2T) in PBD format from the protein data bank was accomplished [48]. Protein Preparation Wizard of Schrödinger-Maestro v9.4 was used for the preparation and refinement of the downloaded protein [49]. Charges and bond orders were assigned and water molecules were deleted. Hydrogens were added to the heavy atoms. Energy minimization was done by using OPLS 2005 force field by fixing the heavy atom RMSD of 0.30Å [50]. Amino acids were optimized by using neutral pH.
2.2 Ligand Preparation
From the Pubchem database the 3D structure of the apigetrin (Pubchem ID; CID 5280704; Molecular formula; C21H20O10) and rutin (Pubchem ID; CID 5280805; Molecular formula; C27H30O16) was downloaded. Ligand preparation was done to create three dimensional geometries and to assign proper bond orders [51]. Three dimensional geometries were generated by using Ligprep2.5 in Schrödinger Suite 2013 with an OPLS_2005 force field [50]. For the generation of ionization states, we used Epik2.2 in Schrödinger Suite at pH7.0±2.0 [52]. A maximum of 32 possible stereoisomers per ligand were obtained.
2.3 Receptor grid generation
During docking trajectory the every poses binds to the predicted active site that’s why receptor grids were calculated for the prepared protein. For Glide docking, grids were generated by using OLPS 2001 force field by keeping the van der Waals scaling factor of 1.0 and charge cutoff value of 0.25. A box was generated to each direction with 14 Å × 14 Å × 14 Å for docking experiments.
2.4 Extra Precision (XP) ligand docking
XP ligand docking was performed rather than SP docking because XP is better than SP in scoring function and it also predicts the false positive results [53]. This docking was performed in Glide of Schrödinger-Maestro v9.4 [54]. Final result of docking can be found as glide score by energy minimization. For docking, van der Waals scaling factor was set to 0.85 and 0.15 for ligand compounds and partial charges cutoff value was fixed at -10.0 kcal/mole. The lowest glide score containing compounds were then subjected to MM-GBSA analysis for binding free energy calculation and best poses were recorded for every ligand compounds.
2.5 Prime MM-GBSA
Binding free energy calculation was also carried out for the protein ligand complexes. MM-GBSA is a combined method for binding free energy calculation which was used in this experiment that accumulates OPLSAA molecular mechanics energies (EMM), an SGB solvation model for polar solvation (GSGB), and a non-polar solvation term (GNP) composed of the non-polar solvent accessible surface area and van der Waals interactions [55]. The best poses from the Glide score were used for binding free energy calculation. The total free energy of binding:
ΔGbind = Gcomplex – (Gprotein + Gligand ), where G = EMM + GSGB + GNP
2.6 Ligand based ADME analysis
For the analysis of physiological descriptors of a compound such as adsorption, distribution, metabolism and excretion behavior of the ligand compounds ADME analysis was done in QikProp module of Schrodinger [56]. It also predicts the physicochemical nature of the compounds as well as their pharmacokinetics properties. In this study, we used the Qikprop 3.2 module of Schrodinger [57]. There are also several other descriptors also analyzed such as Predicted IC50 for blocking HERG K+ channel in vitro, predicted octanol or water partition coefficient [log P(o/w)], number of hydrogen bond acceptors (HBA), number of hydrogen bond donors (HBD), predicted aqueous solubility (log s), solvent-accessible surface area (SASA), skin permeability (log Kp), MDCK cell permeability (MDCK), binding to human serum albumin (log Khsa), blood-brain partition coefficient (logBB), percentage human oral absorption rate.
3.1 Molecular Docking analysis
The docking study of the two ligand compounds revealed that the compound rutin had the highest docking score and another compound apigetrin exhibited much lower docking score than rutin (Table 1). For residue interactions of ligand compounds with the protein molecule, we analyzed the protein-ligand complex structures and found that;
Table 1: Molecular docking results of rutin and apigetrin in kcal/mol
|
Compound Name |
ΔGbind |
Docking score |
Glide energy |
Glide ligand efficiency |
Strain penalty |
Glide Emodel |
|
Rutin |
-44.044 |
-9.618 |
-57.033 |
-0.223 |
0 |
-83.133 |
|
Apigetrin |
-47.351 |
-4.808 |
-44.244 |
-0.155 |
0 |
-58.087 |
The first compound rutin (Figure 2) formed seven conventional hydrogen bonds with Gly69, Glu71, Asn61, His44, Arg48, Ser29 and Asp123 residues where Arg48 made double hydrogen bonds. Glu71 also made double bonds while Pi-alkyl bond was observed for Val31. Trp68 made one pi-pi T stacked bond.
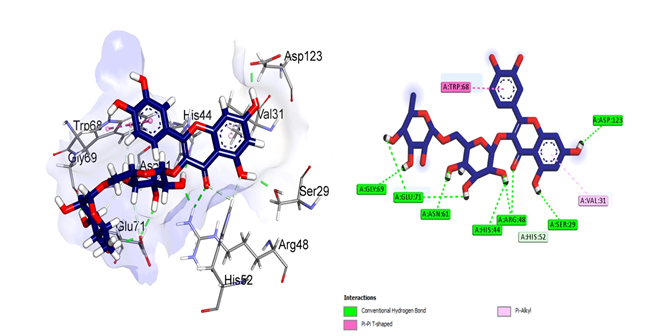
Figure 2: Structural representation of Molecular Docking analysis of rutin with human galectin-1 protein. Rutin binds to the residues of CRD domain and block the active site (The image on the left side give three dimensional overview of the interaction of rutin with human galectin-1 protein and the image on the right side explain the two dimensional binding pattern of rutin with galectin-1)
The second compound apigetrin (Figure 3), on the other hand, made five hydrogen bonds such as Asn33, Lys63, Glu71, Tyr119 and Asp123. All of them showed single bond except Tyr119 who made two conventional hydrogen bonds. His44 made two pi-pi stacked bonds while Val31 made two pi-alkyl bonds.
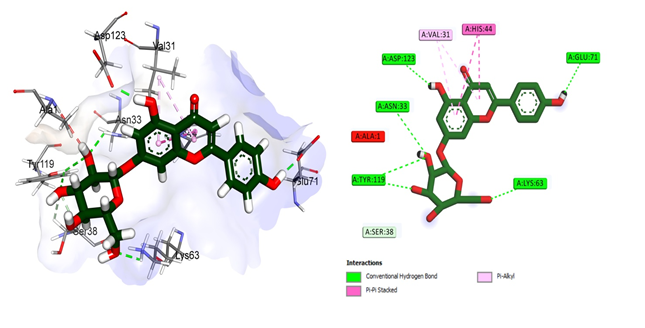
Figure 3: Structural representation of Molecular docking analysis of apigetrin with human galectin-1 protein. Apigetrin made interactions with the residues of CRD domain and block the active site (The image on the left side give three dimensional overview of the interaction of apigetrin with human galectin-1 protein and the image on the right side explain the two dimensional binding pattern of apigetrin with galectin-1)
From the previously published crystal structure of the Human galectin-1 protein it was found that the protein has 10 residues in CRD domain which consists of His52, Arg48, Asn46, Ser29, Asp123, Val31, His44, Asn33, Glu71 and Asn61 [48]. For the stabilization of ligand compound with the protein molecule, interaction should be made between His44, Asn46, Arg48, His52, Asn61, Trp68, Glu71 and Arg73 residues [6]. From our analysis, we found that the targeted ligands were able to make contact with nearly all of the residues in the CRD domain. The interactions made by corresponded ligands with Val31, Ser29, Arg48, His44, Asn33, Glu71, Asn61 and Asp123 residues imply that they can successfully block the CRD domain of the protein.
In another study, human galectin-1 protein in complex with thiodigalactoside (TDG) revealed that, the disaccharide TDG, made hydrogen bond with Glu71 and pi-stacked bond with Trp68, which evidently supports our study [58]. The targeted ligands in this study also made hydrogen bond with Glu71 and pi-stacked bond with Trp68. However, TDG made two additional hydrogen bonds with Asp54 and Arg73 where Asp54 is not involved in the active site of the CRD domain. Interestingly, the ligands of this study, made hydrogen bonds with Asn61, His44, Arg48, Ser29, Asp123 and Asn33. These residues were involved in ligand binding efficiency of CRD domain. Thus this study showed greater binding efficiency of ligands toward the galectin-1 protein than the previous study.
Another experiment, involved interactions between galetin-1 protein with thiodigalactoside derivative, TDG139, where interactions were made by Ser29, Val31, Asp54, Arg73, Glu71, Arg48 and Asn46 residues [59]. Ligands of this experiment made common interactions by Ser29, Val31, Arg48 and Glu71 residues, which were also supported by previous study. A study was carried out to investigate the interaction pattern of galetin-1 protein with type 1 N-acetyllactosamine [60]. Hydrogen bonds were observed for His44, His52, Arg48, Arg73, Trp68 and Asp54 residues. Trp68 also made van der Waals interaction. In this study, interactions were analyzed for three residues in common such as Arg48, His44 and Trp68 residues.
Overall, this study shared most of the residues that were investigated to be involved in the interactions between galectin-1 protein and other published ligand complexes thus indicating the potentiality of rutin and apigetrin as galectin-1 inhibitor.
3.2 Binding free energy calculation
According to the MM-GBSA analysis it was found that, the compound rutin had a binding free energy of -44.04 kcal/mol, whereas the compound apigetrin exhibited a binding free energy of -47.35 kcal/mol (Table 1). From the result, it can be easily observed that, apigetrin has the greater binding free energy than rutin thus showed stronger binding than rutin. In case of coulomb energy, rutin showed the value of -30.44 kcal/mol and apigetrin showed -29.44 kcal/mol. The packing energy released for rutin ligand was less (-4.24 kcal/mol) than the packing energy of apigetrin (-7.12 kcal/mol). Energy released during van der Waals interaction was also lower in case of rutin which was -28.68 kcal/mol and for apigetrin it was -35.00 kcal/mol. From the MM-GBSA analysis it was found that, the apigetrin exhibited higher binding free energy than rutin which has stronger affinity for binding with human galectine-1 protein than rutin. Monosaccharide (Apigetrin) showed better ligand binding efficiency than disaccharide (Rutin) according to the binding free energy calculation of this study.
3.3 ADME analysis
The ADME properties of both rutin and apigetrin were evaluated to explain their pharmacokinetic properties. Table 2 illustrates ADME properties of these two compounds. The properties represent the bioavailability, distribution, cell permeability, excretion and absorption quality of the compounds. From the results of ADME analysis, it was observed that, the blood or brain barrier permeability of the tested compounds was nearly between the acceptable ranges which is very important for a drug to pass through those barriers. Apigetrin showed QPlogBB value of -3.151 which is a better than rutin (-4.728) where the acceptable range is -3.0 to 1.2. The number of hydrogen bonds donor and acceptor are in the value of acceptable range and solvent-accessible surface area (SASA) also showed acceptable value. Predicted IC50 value for blocking HERG K+ channel was very close to the acceptable range for both rutin (-5.288) and apigetrin (-5.799). The predicted octanol or water partition coefficient for rutin and apigetrin were also analyzed. Apigetrin showed better result than rutin by providing the acceptable value of -0.307 where Rutin showed -2.57 value where the acceptable range is -2.0 to 6.5. Human oral absorption rate was also greater for apigetrin (30%) than rutin (0%) according to the findings of this study. In case of cell permeability, apigetrin again showed better result than rutin, which is a very important parameter for a drug to pass through the cell to be active. Skin permeability was near to the acceptable range for both rutin and apigetrin.
Table 2: ADME properties of compounds using Qikprop
|
Name |
QPlogaBB |
HBb donor |
HBc acceptor |
SASAd |
QP log HERGe |
QP log Sf |
QP log Po/wg |
% Human Oral Absorptionh |
QPPMDCKi |
QPlogKpj |
|
Rutin |
-4.728 |
9 |
20.55 |
802.165 |
-5.288 |
-2.284 |
-2.57 |
0 |
0.19 |
-7.581 |
|
Apigetrin |
-3.151 |
5 |
12.25 |
680.454 |
-5.799 |
-3.248 |
-0.307 |
30.657 |
3.69 |
-5.528 |
aPredicted blood/brain partition coefficient, QPlogBB = -3.0-1.2
bHydrogen bonds donor, HB donor = 0.0-6.0
cHydrogen bonds acceptor, HB acceptor = 2.0-20.0
dTotal solvent accessible surface area, SASA = 300.0-1000.0
ePredicted IC50 value for blockage of HERG K+ channels, QPlogHERG = Concern below -5
f Predicted aqueous solubility, QPlogS = -6.5-0.5
gPredicted octanol/water partition coefficient, QP log Po/w = -2.0-6.5
hPredicted qualitative human oral absorption, (%) = >80% is higher, <25% is poor
iPredicted apparent MDCK cell permeability in nm/sec, QPPMDCK= >500 is great, <25 is poor
jPredicted skin permeability, QPlogKp = -8.0 to -10.0
Human galectin-1 is now become a promising target to treat cancer. According to our study, the apigetrin and rutin occupied the CRD domain of the galectin-1 by interacting with most of the residues of CRD domain such as Val31, Ser29, Arg48, His44, Asn33, Glu71, Asn61 and Asp123 which were also supported by previous studies thus can inhibit the galectin-1. Apigetrin also showed 30% of human oral consumption rate which is in medium level quality as well as exhibit better quality as a drug than rutin in most of the parameters of ADME analysis. Thus further in vivo study is needed to evaluate the quality of the apigetrin as a drug compound and its inhibitory activity.
Drickamer, K., Two distinct classes of carbohydrate-recognition domains in animal lectins. J. biol. Chem, 1988. 263(20): p. 9557-9560. PMid:3290208
PubMed/NCBIHirabayashi, J. and K.-i. Kasai, The family of metazoan metal-independent β-galactoside-binding lectins: structure, function and molecular evolution. Glycobiology, 1993. 3(4): p. 297-304. PMid:8400545 3. Barondes, S.H., et al., Galectins: a family of animal beta-galactoside-binding lectins. Cell, 1994. 76(4): p. 597-8. https://doi.org/10.1016/0092-8674(94)90498-7
View Article PubMed/NCBICamby, I., et al., Galectin-1: a small protein with major functions. Glycobiology, 2006. 16(11): p. 137R-157R. PMid:16840800
View Article PubMed/NCBILeffler, H., et al., Introduction to galectins. Glycoconjugate journal, 2002. 19(7-9): p. 433-440. PMid:14758066
View Article PubMed/NCBILópez-Lucendo, M.F., et al., Growth-regulatory human galectin-1: crystallographic characterisation of the structural changes induced by single-site mutations and their impact on the thermodynamics of ligand binding. Journal of molecular biology, 2004. 343(4): p. 957-970. PMid:15476813
View Article PubMed/NCBIHughes, R.C., Galectins as modulators of cell adhesion. Biochimie, 2001. 83(7): p. 667-676. 01289-5
View ArticleScott, K. and C. Weinberg, Galectin-1: a bifunctional regulator of cellular proliferation. Glycoconjugate journal, 2002. 19(7-9): p. 467-477. PMid:14758070
View Article PubMed/NCBIPerillo, N.L., et al., Apoptosis of T cells mediated by galectin-1. Nature, 1995. 378(6558): p. 736. PMid:7501023
View Article PubMed/NCBIPark, J.W., et al., Association of galectin-1 and galectin-3 with Gemin4 in complexes containing the SMN protein. Nucleic acids research, 2001. 29(17): p. 3595-3602. PMid:11522829
View Article PubMed/NCBIThijssen, V.L., et al., Galectin-1 is essential in tumor angiogenesis and is a target for antiangiogenesis therapy. Proceedings of the National Academy of Sciences, 2006. 103(43): p. 15975-15980. PMid:17043243
View Article PubMed/NCBIMasoura, S., et al., Biomarkers in pre-eclampsia: a novel approach to early detection of the disease. Journal of Obstetrics and Gynaecology, 2012. 32(7): p. 609-616. PMid:22943702
View Article PubMed/NCBIde la Fuente, H., D. Cibrián, and F. Sánchez-Madrid, Immunoregulatory molecules are master regulators of inflammation during the immune response. FEBS letters, 2012. 586(18): p. 2897-2905. PMid:22819828
View Article PubMed/NCBIWada, J. and H. Makino, Galectins, galactoside-binding mammalian lectins: clinical application of multi-functional proteins. Acta Medica Okayama, 2001. 55(1): p. 11-18. PMid:11246972
PubMed/NCBIRabinovich, G., Galectin-1 as a potential cancer target. British journal of cancer, 2005. 92(7): p. 1188. PMid:15785741
View Article PubMed/NCBIBarrow, H., J.M. Rhodes, and L.G. Yu, The role of galectins in colorectal cancer progression. International journal of cancer, 2011. 129(1): p. 1-8. PMid:21520033
View Article PubMed/NCBIDalotto-Moreno, T., et al., Targeting galectin-1 overcomes breast cancer-associated immunosuppression and prevents metastatic disease. Cancer research, 2013. 73(3): p. 1107-1117. PMid:23204230
View Article PubMed/NCBISzöke, T., et al., Prognostic significance of endogenous adhesion/growth-regulatory lectins in lung cancer. Oncology, 2005. 69(2): p. 167-174. PMid:16127288
View Article PubMed/NCBISaussez, S., et al., Galectins as modulators of tumor progression in head and neck squamous cell carcinomas. Head & neck, 2007. 29(9): p. 874-884. PMid:17315170
View Article PubMed/NCBIChow, S., et al., Analysis of protein profiles in human epithelial ovarian cancer tissues by proteomic technology. European journal of gynaecological oncology, 2010. 31(1): p. 55-62. PMid:20349782
PubMed/NCBILaderach, D.J., et al., A unique galectin signature in human prostate cancer progression suggests galectin-1 as a key target for treatment of advanced disease. Cancer research, 2013. 73(1): p. 86-96. PMid:23108139
View Article PubMed/NCBIRorive, S., et al., Galectin‐1 is highly expressed in human gliomas with relevance for modulation of invasion of tumor astrocytes into the brain parenchyma. Glia, 2001. 33(3): p. 241-255. 33:3<241::AID-GLIA1023>3.0.CO;2-1
View ArticleCroci, D.O., et al., Disrupting galectin-1 interactions with N-glycans suppresses hypoxia-driven angiogenesis and tumorigenesis in Kaposi's sarcoma. Journal of Experimental Medicine, 2012. 209(11): p. 1985-2000. PMid:23027923
View Article PubMed/NCBIKoopmans, S.M., et al., The involvement of Galectins in the modulation of the JAK/STAT pathway in myeloproliferative neoplasia. American journal of blood research, 2012. 2(2): p. 119. PMid:22762031
PubMed/NCBID'Haene, N., et al., The differential expression of Galectin-1 and Galectin-3 in normal lymphoid tissue and non-Hodgkin's and Hodgkin's lymphomas. International journal of immunopathology and pharmacology, 2005. 18(3): p. 431-443. PMid:16164826
View Article PubMed/NCBIThijssen, V.L., et al., Tumor cells secrete galectin-1 to enhance endothelial cell activity. Cancer research, 2010. 70(15): p. 6216-6224. PMid:20647324
View Article PubMed/NCBIHsieh, S., et al., Galectin-1, a novel ligand of neuropilin-1, activates VEGFR-2 signaling and modulates the migration of vascular endothelial cells. Oncogene, 2008. 27(26): p. 3746. PMid:18223683
View Article PubMed/NCBIMercier, M.L., et al., Knocking down galectin 1 in human hs683 glioblastoma cells impairs both angiogenesis and endoplasmic reticulum stress responses. Journal of Neuropathology & Experimental Neurology, 2008. 67(5): p. 456-469. PMid:18431251
View Article PubMed/NCBIRabinovich, G.A., et al., Synthetic lactulose amines: novel class of anticancer agents that induce tumor-cell apoptosis and inhibit galectin-mediated homotypic cell aggregation and endothelial cell morphogenesis. Glycobiology, 2005. 16(3): p. 210-220. PMid:16282605
View Article PubMed/NCBIIurisci, I., et al., Synthetic inhibitors of galectin-1 and-3 selectively modulate homotypic cell aggregation and tumor cell apoptosis. Anticancer research, 2009. 29(1): p. 403-410. PMid:19331179
PubMed/NCBIMiller, M.C., A. Klyosov, and K.H. Mayo, The α-galactomannan Davanat binds galectin-1 at a site different from the conventional galectin carbohydrate binding domain. Glycobiology, 2009. 19(9): p. 1034-1045. PMid:19541770
View Article PubMed/NCBIPoirier, F., et al., Expression of the L14 lectin during mouse embryogenesis suggests multiple roles during pre-and post-implantation development. Development, 1992. 115(1): p. 143-155. PMid:1638977
PubMed/NCBIIto, K. and S.J. Ralph, Inhibiting galectin-1 reduces murine lung metastasis with increased CD4+ and CD8+ T cells and reduced cancer cell adherence. Clinical & experimental metastasis, 2012. 29(6): p. 561-572. PMid:22484915
View Article PubMed/NCBIDelaine, T., et al., Galectin-inhibitory thiodigalactoside ester derivatives have antimigratory effects in cultured lung and prostate cancer cells. Journal of medicinal chemistry, 2008. 51(24): p. 8109-8114. PMid:19053747
View Article PubMed/NCBIIto, K., et al., Thiodigalactoside inhibits murine cancers by concurrently blocking effects of galectin-1 on immune dysregulation, angiogenesis and protection against oxidative stress. Angiogenesis, 2011. 14(3): p. 293-307. PMid:21523436
View Article PubMed/NCBIAstorgues-Xerri, L., et al., Unraveling galectin-1 as a novel therapeutic target for cancer. Cancer treatment reviews, 2014. 40(2): p. 307-319. PMid:23953240
View Article PubMed/NCBIMagedov, I.V., et al., Discovery and investigation of antiproliferative and apoptosis-inducing properties of new heterocyclic podophyllotoxin analogues accessible by a one-step multicomponent synthesis. Journal of medicinal chemistry, 2007. 50(21): p. 5183-5192. PMid:17894480
View Article PubMed/NCBIGordaliza, M., Natural products as leads to anticancer drugs. Clinical and Translational Oncology, 2007. 9(12): p. 767-776. PMid:18158980
View Article PubMed/NCBIRollinger, J., T. Langer, and H. Stuppner, Strategies for efficient lead structure discovery from natural products. Current medicinal chemistry, 2006. 13(13): p. 1491-1507. PMid:16787200
View Article PubMed/NCBIAmin, A.R., et al., Perspectives for cancer prevention with natural compounds. Journal of clinical oncology, 2009. 27(16): p. 2712. PMid:19414669
View Article PubMed/NCBIMurakami, A., H. Ashida, and J. Terao, Multitargeted cancer prevention by quercetin. Cancer letters, 2008. 269(2): p. 315-325. PMid:18467024
View Article PubMed/NCBIDuthie, S.J. and V. Dobson, Dietary flavonoids protect human colonocyte DNA from oxidative attack in vitro. European Journal of Nutrition, 1999. 38(1): p. 28-34. PMid:10338685
View Article PubMed/NCBIDeschner, E.E., et al., Quercetin and rutin as inhibitors of azoxymethanol-induced colonic neoplasia. Carcinogenesis, 1991. 12(7): p. 1193-1196. PMid:2070483
View Article PubMed/NCBISrivastava, J.K. and S. Gupta, Extraction, characterization, stability and biological activity of flavonoids isolated from chamomile flowers. Molecular and cellular pharmacology, 2009. 1(3): p. 138. PMid:20098626
View Article PubMed/NCBITejler, J., et al., Fragment-based development of triazole-substituted O-galactosyl aldoximes with fragment-induced affinity and selectivity for galectin-3. Organic & biomolecular chemistry, 2009. 7(19): p. 3982-3990. PMid:19763301
View Article PubMed/NCBIvan Hattum, H., et al., Tuning the preference of thiodigalactoside-and lactosamine-based ligands to galectin-3 over galectin-1. Journal of medicinal chemistry, 2013. 56(3): p. 1350-1354. PMid:23281927
View Article PubMed/NCBIBlanchard, H., et al., Galectin-1 inhibitors and their potential therapeutic applications: a patent review. Expert opinion on therapeutic patents, 2016. 26(5): p. 537-554. PMid:26950805
View Article PubMed/NCBICollins, P.M., et al., Taloside Inhibitors of Galectin‐1 and Galectin‐3. Chemical biology & drug design, 2012. 79(3): p. 339-346. PMid:22136701
View Article PubMed/NCBIDash, R., et al., In silico analysis of indole-3-carbinol and its metabolite DIM as EGFR tyrosine kinase inhibitors in platinum resistant ovarian cancer vis a vis ADME/T property analysis. 2015.
Shivakumar, D., et al., Prediction of absolute solvation free energies using molecular dynamics free energy perturbation and the OPLS force field. Journal of chemical theory and computation, 2010. 6(5): p. 1509-1519. PMid:26615687
View Article PubMed/NCBISastry, G.M., et al., Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. Journal of computer-aided molecular design, 2013. 27(3): p. 221-234. PMid:23579614
View Article PubMed/NCBIWizard, P.P., Epik version 2.2, Impact version 5.7, Prime version 3. New York, NY: Schrödinger, LLC, 2011.
Friesner, R.A., et al., Extra precision glide: Docking and scoring incorporating a model of hydrophobic enclosure for protein− ligand complexes. Journal of medicinal chemistry, 2006. 49(21): p. 6177-6196. PMid:17034125
View Article PubMed/NCBIFriesner, R.A., et al., Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. Journal of medicinal chemistry, 2004. 47(7): p. 1739-1749. PMid:15027865
View Article PubMed/NCBIRastelli, G., et al., Fast and accurate predictions of binding free energies using MM‐PBSA and MM‐GBSA. Journal of computational chemistry, 2010. 31(4): p. 797-810. PMid:19569205
PubMed/NCBINatarajan, A., et al., Molecular docking studies of (4 Z, 12 Z)-cyclopentadeca-4, 12-dienone from Grewia hirsuta with some targets related to type 2 diabetes. BMC complementary and alternative medicine, 2015. 15(1): p. 73. PMid:25885803
View Article PubMed/NCBISharma, V., et al., Structure based rational drug design of selective phosphodiesterase-4 ligands as anti-inflammatory molecules. Bull. Pharm. Res, 2011. 1(2): p. 33-40.
Stannard, K.A., et al., Galectin inhibitory disaccharides promote tumour immunity in a breast cancer model. Cancer letters, 2010. 299(2): p. 95-110. PMid:20826047
View Article PubMed/NCBIHsieh, T.-J., et al., Dual thio-digalactoside-binding modes of human galectins as the structural basis for the design of potent and selective inhibitors. Scientific reports, 2016. 6: p. 29457. PMid:27416897
View Article PubMed/NCBIHsieh, T.-J., et al., Structural basis underlying the binding preference of human galectins-1,-3 and-7 for Galβ1-3/4GlcNAc. PloS one, 2015. 10(5): p. e0125946. PMid:25945972
View Article PubMed/NCBI