Hongfeng Bian
E-mail:bianhf108@nenu.edu.cn
© 2019 Sift Desk Journals. All Rights Reserved
VOLUME: 5 ISSUE: 2
Page No: 74-82
Hongfeng Bian
E-mail:bianhf108@nenu.edu.cn
1School of Environment, Northeast Normal University,ChangChun 130024, China
Shuangshuang Wu, Min Xu, Jianwei Li, Hongfeng Bian, Bacterial community structure and soil enzyme activities response to change of moisture content in northeastern China forest peat swamps (2020) Journal of Earth Sciences & Environmental Studies 5(2) pp:74-82
The better understanding of the relationship between soil physical and chemical properties and bacterial communities under different moisture contents has a potential contribution to improving the sustainability of forest peat swamp protection. In this study, we tested the 16sRNA genes of bacteria to study the physical and chemical characteristics of soil under three types of forest surface vegetation cover, with three different moisture contents (24.4 ± 2.6%, 48.4 ± 4.2%, 91.0 ± 2.7%). The results show that under different moisture contents conditions, microbial community composition will change greatly. Proteobacteria (30.04%) and Acidobacteria (25.58%) are the dominant communities in forest peat swamps, and have little change in moisture contents , Chloroflexi abundance changes significantly in different moisture contents, the microbial community structure has a strong selectivity to water. As the moisture content of the forest peat swamp decreases, the soil microbial community structure will change, and soil enzyme activities (Invertase, Urease, Catalase) have different activities under different water contents, which impact on deposition rate of C in the soil and the carbon sequestration of organic matter in forest peat swamps.
Keywords: forest swamp; soil enzyme activity; bacterial community; global warming; soil moisture content
Northern forests account for 22% of the global forest area. These small forests account for only 13% of global biomass carbon. Their peat-rich soil account for nearly half (43%) of the global peat reserve, which is equivalent to 455 Pg carbon pool1,2. The northern peat bog is a transition zone between terrestrial aquatic ecosystems3. It has high resource development value and environmental regulation functions, and has a lasting positive response to the global climate system2,4. With the change of climate, the frequency and intensity of drought are expected to increase in the 21st century5. As a transitional zone from aquatic to terrestrial, wetlands are more susceptible to hydrological changes under the trend of global warming6. The impact of human activities has made the drying of wetlands increasingly apparent. Under the background of global warming, the evaporation of forest peat swamp wetlands has risen, affecting vegetation coverage in turn affecting soil enzyme activity and microbial distribution7. Soil microorganisms, including protozoa, bacteria, fungi, and archaea, are important constituents of the soil. They play an important role in the biogeochemical cycle, especially in the processes of carbon cycling, ecosystem stability, sustainable soil development and anti-interference ability8. The soil bacterial community structure is critical to the function of the wetland system, and they mediate the material cycle in the environment8. Many studies have focused on the spatial and temporal differences in the structure of soil bacterial communities9. Forest peat swamps are important global ecosystems, and little known about their ecological importance and biodiversity10. Northeast China is one of the most important peat distribution areas in the world, but few studies have focused on the relationship between the bacterial community structure in forest peat swamps and the physical and chemical properties of soil in peat wetlands11–13.
The activity of microorganisms and enzymes usually changes with temperature, humidity, pH and other physical and chemical variables, as well as the utilization of carbon and nutrients7. Alien microorganisms and their impacts on mineralization of organic matter and the global carbon and nutrient cycle play a decisive role8,11. The most direct mediator of soil organic matter (SOM) decomposition is extracellular enzymes9. The presence of extracellular enzymes directly deconstructs the cell walls of plants and microorganisms10. And the macromolecules are reduced to soluble substrates, to assimilate with microorganisms. Therefore, studying the metabolic relationship between microorganisms and enzymes in northern forest peat swamps has an important role in guiding the protection of forest peat swamps12. More and more scholars have begun to pay attention to the problems of microbial and enzyme activities in peat swamps13. The importance of changes in the composition of soil microbial communities to the ecosystem depends on whether environmental changes will affect changes in soil nutrients and carbon cycling in the soil14–17. Soil enzyme activity also has a corresponding change in the microbial community structure in the soil, so we also measured the enzyme activity in the soil to determine whether the composition of the microbial community could further explain the migration of soil carbon under different hydrological gradients18,19.
The purpose of this study was to study the bacterial community structure and soil physicochemical properties of soil in three different moisture content rates in two typical forest-swamp wetlands in northeastern China. We hypothesized that different moisture content will cause significant changes in the soil microbial community structure; the main reason for the difference in soil microbial community is the soil water content, and the enzyme activity is mainly regulated by soil nutrients; There is a relationship between structural changes.
2.1 In-situ sample collection
Due to the strong metabolic activity of microorganisms in summer, the experiment was conducted at the end of May 2019. The experimental research area is in Longwan Nature Reserve in Tonghua City, Jilin Province. Two typical secondary forest ecosystems in this area were selected for research20. The area is located at the northern foot of Changbai Mountain, where is the middle section of the Longgang Mountains belongs to a moderate-temperate continental monsoon climate, with an average annual temperature of 4-5 ° C and an average annual rainfall of 800-900mm. Gushantun (N 42 ° 18'36.5076 ", E 126 ° 17'38.3856", 514m above sea level) Vegetation types are mainly white birch (Betula platyphyla) and poplar(Populus davidiana),Hani (N 42 ° 8'24.3312 ", E 126 ° 18'54.7416", 912m above sea level) The vegetation type is dominated by deciduous pine forest (CBS). The forest plots in the study area are mainly dark brown forest soil, and valley valleys are mainly meadow soils21. Plateau lessive soil (white pulp soil) is dominant. Changbai Mountain is in the temperate coniferous forest-broadleaf forest belt and is one of the best-preserved virgin forests in China. In this experiment, six sampling areas were selected for analysis in three types of forest marshes with different soil moisture content. A sample plot of 10m ⅹ 10m was set, and three trees with substantially the same DBH were randomly selected in the plot, and 30cm from the trunk Take a soil sample at a depth of 0-10cm. At each sampling point, biomass was removed from the ground and soil samples were evenly mixed. The recovered soil sample was divided into two parts after removing visible roots and residues as soon as possible. One sample was air-dried and ground into a fine powder and stored through a 100-mesh sieve for enzyme activity and soil physical and chemical properties determination. Another sample was stored at -80 ° C until high-throughput sequencing.
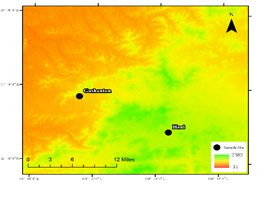
Figure 1
2.2 Analysis of soil physical and chemical properties
The soil pH was measured at a soil-to-water ratio of 1: 2.5. The total carbon and total nitrogen (TC, TN) in the soil were determined using a C / N analyzer. Since these soils without carbonates, the total carbon content is equal to the organic carbon (SOC) content in the soil. The available phosphorus (AP) is determined by the molybdenum antimony phosphate colorimetric method. Soil nitrate nitrogen (NO3-N) and ammonium nitrogen (NH4+ -N) are extracted from 5g fresh soil ) and analyzed in 2MKCl (soil: extractant = 1: 5) using a flow injection automatic analyzer.
Enzyme activity in soil, in short for invertase activity, mix 5 g soil with 15 ml 8% sucrose, 5 ml of phosphate buffer pH 5.5 and 5 drops of toluene, then incubate at 37 ° C for 24 hours22. The released glucose was reacted with 3,5-dinitrosalicylic acid and measured by absorption spectrophotometry at 508 nm. For urease activity, 5g of fresh bulk soil (<2mm) was incubated with 10mL of 10% urea, 1mL of toluene and 20mL of PB buffer (pH 6.7) at 37°C for 24h23. The absorption spectrophotometry of indophenol at 578 nm was measured. For catalase, 5g of fresh soil (<2mm) and 40mL of distilled water and 5mL of 0.3% hydrogen peroxide solution were shaken for 20min, and 5mL of 2mol / L sulfuric acid solution was added to terminate the enzyme reaction24. The potassium permanganate solution was used to titrate the hydrogen peroxide remaining after the catalase decomposition reaction.
2.3 DNA Extraction and 16S rRNA Sequencing and Sequence Processing
Bacterial communities on soil were detected by next-generation amplicon sequencing of 16S rRNA genes. PCR was applied to amplify the V3-V4 hypervariable region of bacterial and archaeal 16S rRNA genes by using the forward primer, 5'-CTTGGTCATTTAGAGGAAGTAA -3' and the reverse primer, 5'- GGACTACHVGGGTWTCTAAT-3'. combined with adapter sequences and barcode PCR amplification was performed in a total volume of 50 μl, which contained 10 μl Bμffer, 0.2 μl Q5 High-Fidelity DNA Polymerase, 10 μl High GC Enhancer, 1 μl dNTP, 10μM of Each primer and 60 ng genome DNA25,26. Thermal cycling conditions were as follows: an initial Denaturation at 95 °C for 5 min, followed by 15 cycles at 95 °C for 1 min, 50 °C for 1 min and 72 °C for 1 min, with a final extension at 72 °C for 7 min. The PCR products from the first step PCR was purified through VAHTSTM DNA Clean Beads. A second round PCR was then Performed in a 40μl reaction which contained 20 μl 2×Phμsion HF MM, 8 μl ddH2O, 10μM of Each primer and 10μl PCR products from the first step. Thermal cycling conditions were as Following: an initial denaturation at 98 °C for 30s, followed by 10 cycles at 98 °C for 10s, 65 °C for 30s min and 72 °C for 30s, with a final extension at 72 °C for 5 min. Finally, all PCR products Were quantified by Quant-iTTM dsDNA HS Reagent and pooled together. High-throughput Sequencing analysis of bacterial rRNA genes was performed on the purified, pooled sample using The Illumina Hiseq 2500 platform (2×250 paired ends) at Biomarker Technologies Corporation, Beijing, China.
2.4 Data Analysis and Statistics
Separate the data for each sample from the original data according to standard operating procedures based on the barcode and primer sequences27. Vsearch v2.4.4 is used to stitch the readings of each sample to obtain raw label data (raw labels). Sequences shorter than 150 bp and of low quality (quality score <20) were deleted from the original sequence data, and a classification operation unit (OUT) was defined according to the similarity between the sequences above 97%.Based on the SILVA128 database, annotate representative OTU sequences with species, use QIIME software to estimate Alpha diversity for Chao 1, Simpson, and Shannon indices, and compare the OTU abundance and uniformity between samples, and then generate dilution curves. The principal component analysis (PCoA) based on Bray-Curtis matrix distance was calculated using the Vegan package in R language, and the differences in the bacterial community structure between two different in situ soil hydrological gradients (L, M, H) were compared. Based on the OUT data, Canoco 5.0 was used to explore the relationship between bacterial communities and environmental factors through redundant analysis. The one-way difference analysis (ANOVA) method using the SPSS 22.0 post hoc Tukey HSD test was used to test soil characteristics, various microbial composition ratios, and significant differences in enzyme activity corresponding to different water contents. The image was drawn using Origin 2018 64Bit.
3.1 Soil physical and chemical properties
The soil temperature in the two forest plots varied between 11.1-17.2 ° C. The pH changes between 5.47-6.08, which belongs to slightly acid soil. The soil physicochemical properties of Gushantun broad-leaved forest swamps under different moisture contents are quite different, among which organic matter (SOC) and total nitrogen (TN) are significantly different in broad-leaved forest (p <0.05) and increase with the increase of soil moisture content; In the Hani coniferous forest, the content of each nutrient element is low and there is not much difference between different moisture contents. There was no significant difference in soil nutrient availability (AP NH4-N NO3-N) between the various sites.
Table 1. Soil physical and chemical properties of two forest types with different moisture content
|
|
Gushantun |
Hanni |
||||
|
|
L |
M |
H |
L |
M |
H |
|
pH |
6.08±0.21a |
5.59±0.33ab |
5.47±0.08b |
5.65±0.03a |
5.71±0.09a |
5.81±0.05a |
|
EC(mS/cm) |
0.95±0.31b |
1.14±0.25ab |
1.62±0.21a |
0.5±0.06a |
0.46±0.04a |
0.45±0.13a |
|
SOC((g/kg)) |
38.15±14.06c |
67.77±13.86b |
129.89±2.18a |
37.41±8.82a |
28.36±4.71a |
42.79±12.26a |
|
Available P (mg/kg) |
0.78±0.18a |
0.68±0.37a |
0.82±0.20a |
0.39±0.05b |
0.23±0.04b |
1.21±0.21a |
|
NO3--N(g/kg) |
1.19±0.16a |
0.13±0.06b |
0.48±0.33b |
0.58±0.25a |
0.24±0.03ab |
0.21±0.01b |
|
NH4+-N(g/kg) |
1.78±0.11a |
1.97±0.59a |
2.39±0.49a |
1.75±0.10a |
1.79±0.15a |
1.73±0.34a |
|
T5(℃) |
14.83±0.55a |
10.57±1.29b |
14.53±1.17a |
16.50±0.53a |
17.20±0.3a |
14.53±0.60b |
|
BD(g/cm-3) |
0.70±0.05a |
0.76a |
0.63±0.09a |
0.63±0.03b |
1.04a |
0.55±0.02c |
|
TN(g/kg) |
4.16±0.73b |
7.97±3.09b |
13.20±1.38a |
4.14±1.11a |
2.32±0.67a |
3.23±1.14a |
Note:H:Higher moisture content=91.0±2.7%,M:Medium moisture content=48.4±4.2%,L:Low moisture content=24.4±2.6%。Values indicate means with standard error (n = 3). Significant different water content is indicated by lower case letters (P < 0.05). EC: electrical conductivity.SOC: Soil Organic Carbon. T5℃: Temperature 5cm from the surface. BD:Bulk Density. Letters signify significantly different mean rates as determined by Tukey's HSD test.
3.2 Soil enzyme activity
The responses of enzyme activities to changes in hydrological conditions and forest vegetation cover showed different trends. Invertase activity decreased in two plots with decreasing moisture content. The difference in enzyme activity reached a statistically significant level (p <0.05). Urease did not change significantly in the Hani coniferous forest. The rate increases. It is worth noting that catalase is not sensitive to changes in water gradient in forest swamps, and only small differences can be observed in coniferous forests.
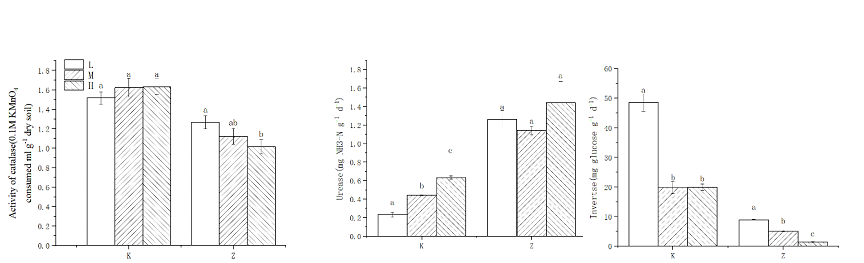
Fig. 2 Changes in different enzyme activities under different moisture contents in the two forest types. Letters signify significantly different mean rates as determined by Tukey's HSD test.
3.3 Influence of environmental factors on the structure of bacterial communities in soil
16S rRNA sequencing identifies the taxonomic composition of bacterial communities. All samples were sequenced to obtain 1,922,134 Reads,Double end Reads splicing and filtering produced a total of 1,787,090 Clean tags. At least 62,926 Clean tags were generated for each sample, and an average of 99,283 Clean tags were generated. According to the dilution curve presented by the sample, it was sufficient. Sequencing depth to explain most of the amplified taxa and ensure the accuracy of sequencing. Based on the analysis of Alpha diversity including Chao1, Shannon, and Simpson index in situ, it is worth noting that the diversity did not show significant differences in the two plots (ANOVA, p> 0.05).
According to the OUT table of bacterial communities, PCoA was used to detect the differences in bacterial communities between different moisture contents. According to PCoA, it can be clearly seen that the bacterial communities on the H plot were significantly different from L and M (PERMANOVA, p <0.05) The bacterial communities in L and M plots were similar, and L and M almost coincided in Hani plots (PERMANOVA, p> 0.05). For the bacterial community species distribution, Proteobacteria, Acidobacteria, Chloroflexi, and Actinobacteria dominate the community in all samples (P <0.05), accounting for 70-80% of the total flora. The abundance of Proteobacteria and Actinobacteria in broadleaf forest marsh soil is greater than that of coniferous forest, while the richness of Chloroflexi in coniferous forest marsh is greater than that of broadleaf forest. The change trend of Chloroflexi with different water content is opposite to that of Proteobacteria.
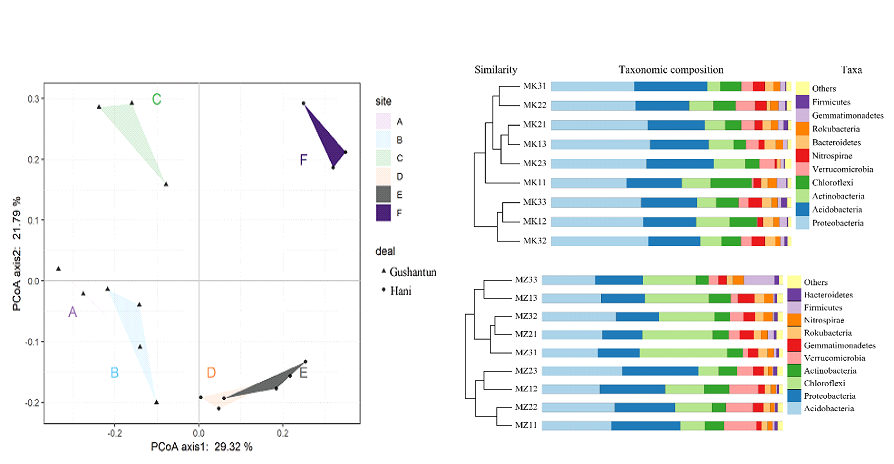
Fig 3. PCoA distribution of the difference index (Bray-Curtis) between bacterial communities. Low and medium moisture content in Gushantun plot: A, medium moisture content: B, high moisture content: C; low and medium moisture content in Hani plot: D, medium moisture content: E, high moisture content: F. Figure 4 Cluster analysis of Hani and Gushantun plots based on the Bray-curtis matrix. Relative abundance maps (n = 3) of bacterial communities (TOP10 of microbial species) at different moisture contents.
The relationship between soil physical and chemical properties and soil microbial characteristics was analyzed by RDA. RDA1 explained 90.49% and 99.37% of the total interpretation rate. It mainly explained the relationship between physical and chemical properties of soil and microbial groups. Controls microbial communities. In Figure 6-1, pH (p = 0.016) and MC (p = 0.024) have significant effects on soil microbial characteristics. In Figure 6-2, MC (p = 0.002) has a significant effect on the sudden microbial community. Compared with other environmental factors, soil moisture is the decisive factor limiting the distribution of microbial communities.
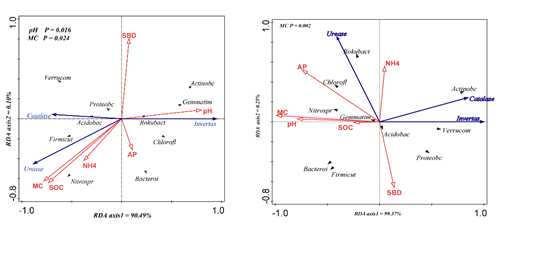
Figure 4.1 Gushantun forest transect
Figure 4.2 Hani forest transect
Figure 4 Redundant analysis (RDA) of soil microorganisms and enzyme activities( Invertase, Urease, Catalase) based on two plots (Gushantun; Hani). Constrained by three hydrological gradients (SBD; pH; AP; NH4; MC; SOC). The first and second axes explain 90.49 and 0.10% in Gushantu forest marsh wetland, respectively. In Hani, the soil interpretation rate is 99.62%, black represents the population of microorganisms, and purple represents soil enzyme activity.
The soil microbial composition between different forest peat swamps showed significant differences. The enzyme activity in the soil was affected by vegetation types, soil nutrients, and soil microorganisms, reflecting changes in soil microbial activity28,29. The role of bacteria in the soil is to maintain sustainable soil development and production, and it plays a huge role in maintaining soil health30. The results of this study can be clearly seen through the changes in soil moisture content of different forest land types. Under different moisture content conditions, the differences in soil enzyme activity and microbial community.
Consistent with our hypothesis, soil moisture is highly correlated with the functional potential of soil microbial communities. Soil microbial communities will be affected by changes in soil moisture, which in turn will affect enzyme activity in the soil. With the exception of catalase, soil enzyme activity appears to be strongly affected by soil moisture content. In the environment with more organic matter (broad-leaved forest), the activity of urease increased with the increase of moisture content, and the activity of invertase decreased in the environment with high moisture content. The transformation of material in the soil is the result of the comprehensive effect of various biochemical reactions in the soil. The strength of soil enzyme activity is one of the important factors of the comprehensive effect of various biochemical reactions in the soil. Invertase is used to catalyze the hydrolysis of sucrose into glucose and fructose, which is related to the content of microorganisms in the soil. In this study, we found that the activity of invertase in soil decreased with the decrease of moisture content, and the soil with low moisture content (23.2%) ,the activity of soil (92.4% MC) with high moisture content is lower than 15.3% -41.0%. The higher the moisture content in the observed soil plot, the higher the invertase activity. Urease is the hydrolysis of urea-based substrates in soil. The main source of urease is the secretion of extracellular microorganisms. We found that urease exhibits different activity trends in two different forest types. In broad-leaved forests, water in the soil can Stimulate the activity of urease (p <0.05). In the RDA ranking chart, we found that urease and other environmental factors except moisture content showed a significant negative correlation. Pearson analysis showed that urease was not significantly related to other environmental factors. Studies have shown that urease is attached to clay and humus and is more resistant to environmental changes than other soil enzymes. Catalase can decompose hydrogen peroxide into water and oxygen molecules and protect cells from being affected by reactive oxygen species. In this study, we found that catalase activity did not change in different forest marsh soil moisture content(P> 0.05). This result is different from Zhang et al. finding that as the moisture content increases, the effect on catalase is more.
The microbial community in the soil drives the decomposition and evolutionary adaptation of the organic matter in the soil. Although the soil moisture content is different, the bacterial community diversity index of the bacterial community in the door has not changed significantly. Comparing the composition of the community structure of the bacteria in the two forests shows that there are obvious differences between the microbial communities in the soils of the broad-leaved forest and the coniferous forest. β diversity is significantly different between bacterial groups. In both plots, the soils with high moisture content showed significantly different soil microbial community distributions than those with low and medium moisture content31. In all the samples, we found that the distribution of bacterial flora was mainly composed of Proteobacteria, Acidobacteria, Chloroflexi, Actinobacteria, Verrucomicrobia, Gemmatimonadetes, Nitrospirae, Rokubacteria, Bacteroidetes, Firmicutes29,32. These floras are resident in the soil. Many studies have shown that plant abundance, soil type, and soil use all affect the community structure of microorganisms33. Gushantun and Hani forest plots have different bacterial community structures. The relative abundance of the dominant phylum Proteobacteria in the Gushantun plot is greater than in the Hani plot34. Chloroflexi changes significantly in abundance in different moisture contents. The results showed that soil moisture mainly affected the supply of soluble carbon and nutrients to soil microorganisms, and soil matrix affected the physiological state of microorganisms. Therefore, the moisture content in soil significantly changed the structure of dominant flora. In soil, microbial communities play an important role in ecosystem functions, biogeochemical functions, and responses to environmental changes. Changes in soil moisture may affect the biomass activity and composition of soil microorganisms35. Soil moisture also remained highly correlated with the microbial community structure in the ranking diagram36–38. Our research found that in the same forest, the difference in soil moisture content caused by the topography did not largely cause changes in the structure of the bacterial community in the soil, which may be due to differences in soil and climate in the same forest community. When it is larger, the vegetation type greatly affects the distribution of microbial communities. Different concentrations of nutrients and different levels of anoxic environment showed significant differences in soil microbial flora distribution and enzyme activity. The microbial community was affected by hydrological conditions such as moisture and oxygen environment in the soil. However, we found that in the forest marsh soil, the biggest impact on the distribution of microbial community structure is not the soil moisture and the conditions of the same period, but the type of vegetation covered by the marsh surface. Over time, microbial communities under different vegetation cover occur has differentiated.
This study shows that in forest peat swamps in northeastern China, the microbial community structure is greatly affected by hydrological gradients. Soil microclimate is an important driving force for the composition of microbial communities. Structure will have a more significant effect, and hydrological conditions have a significant effect on soil enzyme activity. In the context of global warming, the differences in soil moisture content are increasingly significant.
This work was supported by the National Natural Science Foundation of China (NSFC)(No. 41630749).
Ise, T., Dunn, A. L., Wofsy, S. C. & Moorcroft, P. R. High sensitivity of peat decomposition to climate change through water-table feedback. Nat. Geosci. 1, 763-766 (2008).
View ArticleGallego-Sala, A. et al. Latitudinal limits to the predicted increase of the peatland carbon sink with warming. Nat. Clim. Change 8, 907-+ (2018).
Gorham, E. Northern Peatlands - Role in the Carbon-Cycle and Probable Responses to Climatic Warming. Ecol. Appl. 1, 182-195 (1991). PMid:27755660
View Article PubMed/NCBIMatthews, H. D., Eby, M., Ewen, T., Friedlingstein, P. & Hawkins, B. J. What determines the magnitude of carbon cycle-climate feedbacks? Glob. Biogeochem. Cycles 21, GB2012 (2007).
View ArticleKeiser, A. D., Smith, M., Bell, S. & Hofmockel, K. S. Peatland microbial community response to altered climate tempered by nutrient availability. Soil Biol. Biochem. 137, UNSP 107561 (2019).
View ArticleAerts, R. The freezer defrosting: global warming and litter decomposition rates in cold biomes. J. Ecol. 713-724 (2010) doi:10.1111/j.1365-2745.2006.01142.x@10.1111/(ISSN)1365-2745.GLOBWA.
View ArticleDedysh, S. N. Cultivating uncultured bacteria from northern wetlands: knowledge gained and remaining gaps. Front. Microbiol. 2, 184 (2011). PMid:21954394
View Article PubMed/NCBIManzoni, S., Schimel, J. P. & Porporato, A. Responses of soil microbial communities to water stress: results from a meta-analysis. Ecology 93, 930-938 (2012). PMid:22690643
View Article PubMed/NCBIKim, S.-Y., Lee, S.-H., Freeman, C., Fenner, N. & Kang, H. Comparative analysis of soil microbial communities and their responses to the short-term drought in bog, fen, and riparian wetlands. Soil Biol. Biochem. 40, 2874-2880 (2008).
View ArticleMakiranta, P. et al. Indirect regulation of heterotrophic peat soil respiration by water level via microbial community structure and temperature sensitivity. Soil Biol. Biochem. 41, 695-703 (2009).
View ArticleBarcenas-Moreno, G., Gomez-Brandon, M., Rousk, J. & Baath, E. Adaptation of soil microbial communities to temperature: comparison of fungi and bacteria in a laboratory experiment. Glob. Change Biol. 15, 2950-2957 (2009).
View ArticleZhou, L. et al. Soil extracellular enzyme activity and stoichiometry in China's forests. Funct. Ecol. doi:10.1111/1365-2435.13555.
View ArticleMeng, C. et al. Global meta-analysis on the responses of soil extracellular enzyme activities to warming. Sci. Total Environ. 705, 135992 (2020). PMid:31841928
View Article PubMed/NCBITaylor, B. R., Parkinson, D. & Parsons, W. F. J. Nitrogen and Lignin Content as Predictors of Litter Decay Rates: A Microcosm Test. Ecology 70, 97-104 (1989).
View ArticleSinsabaugh, R. L. et al. Stoichiometry of soil enzyme activity at global scale. Ecol. Lett. 11, 1252-1264 (2008). PMid:18823393
View Article PubMed/NCBIJassey, V. E. J. et al. Tipping point in plant-fungal interactions under severe drought causes abrupt rise in peatland ecosystem respiration. Glob. Change Biol. 24, 972-986 (2018). PMid:28991408
View Article PubMed/NCBISong, Y. et al. Linking plant community composition with the soil C pool, N availability and enzyme activity in boreal peatlands of Northeast China. Appl. Soil Ecol. 140, 144-154 (2019).
View ArticleShen, F. et al. Soil N/P and C/P ratio regulate the responses of soil microbial community composition and enzyme activities in a long-term nitrogen loaded Chinese fir forest. Plant Soil 436, 91-107 (2019).
View ArticleWilson, R. M. et al. Stability of peatland carbon to rising temperatures. Nat. Commun. 7, 13723 (2016). PMid:27958276
View Article PubMed/NCBIjha, D. K., Sharma, G. D. & Mishra, R. R. Soil microbial population numbers and enzyme activities in relation to altitude and forest degradation. Soil Biol. Biochem. 24, 761-767 (1992). 90250-2
View ArticleLi, N. et al. Records of East Asian monsoon activities in Northeastern China since 15.6 ka, based on grain size analysis of peaty sediments in the Changbai Mountains. Quat. Int. 447, 158-169 (2017).
View ArticleTabatabai, M. A. Soil Enzymes. in Encyclopedia of Agrochemicals (American Cancer Society, 2003). doi:10.1002/047126363X.agr354.
View ArticleMarschner, B. & Kalbitz, K. Controls of bioavailability and biodegradability of dissolved organic matter in soils. Geoderma 113, 211-235 (2003). 00362-2
View ArticleChen, H. et al. Reduced tillage and increased residue retention increase enzyme activity and carbon and nitrogen concentrations in soil particle size fractions in a long-term field experiment on Loess Plateau in China. Soil Tillage Res. 194, 104296 (2019).
View ArticleGardes, M. & Bruns, T. D. ITS primers with enhanced specificity for basidiomycetes - application to the identification of mycorrhizae and rusts. Mol. Ecol. 2, 113-118 (1993). PMid:8180733
View Article PubMed/NCBIŠrut, M., Menke, S., Höckner, M. & Sommer, S. Earthworms and cadmium - Heavy metal resistant gut bacteria as indicators for heavy metal pollution in soils? Ecotoxicol. Environ. Saf. 171, 843-853 (2019). PMid:30660978
View Article PubMed/NCBIQuast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590-D596 (2013). PMid:23193283
View Article PubMed/NCBIBakker, M. G., Otto-Hanson, L., Lange, A. J., Bradeen, J. M. & Kinkel, L. L. Plant monocultures produce more antagonistic soil Streptomyces communities than high-diversity plant communities. Soil Biol. Biochem. 65, 304-312 (2013).
View ArticleHelfrich, M., Ludwig, B., Thoms, C., Gleixner, G. & Flessa, H. The role of soil fungi and bacteria in plant litter decomposition and macroaggregate formation determined using phospholipid fatty acids. Appl. Soil Ecol. 96, 261-264 (2015).
View ArticleRichards, A. E., Forrester, D. I., Bauhus, J. & Scherer-Lorenzen, M. The influence of mixed tree plantations on the nutrition of individual species: a review. Tree Physiol. 30, 1192-1208 (2010). PMid:20472645
View Article PubMed/NCBICaporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. Isme J. 6, 1621-1624 (2012). PMid:22402401
View Article PubMed/NCBIHaettenschwiler, S., Aeschlimann, B., Couteaux, M.-M., Roy, J. & Bonal, D. High variation in foliage and leaf litter chemistry among 45 tree species of a neotropical rainforest community. New Phytol. 179, 165-175 (2008). PMid:18422903
View Article PubMed/NCBIGuo, Q. et al. Species-specific competition and N fertilization regulate non-structural carbohydrate contents in two Larix species. For. Ecol. Manag. 364, 60-69 (2016).
View ArticleChung, H., Zak, D. R., Reich, P. B. & Ellsworth, D. S. Plant species richness, elevated CO2, and atmospheric nitrogen deposition alter soil microbial community composition and function. Glob. Change Biol. 13, 980-989 (2007).
View ArticleDrenovsky, R. E., Vo, D., Graham, K. J. & Scow, K. M. Soil water content and organic carbon availability are major determinants of soil microbial community composition. Microb. Ecol. 48, 424-430 (2004). PMid:15692862
View Article PubMed/NCBIJangid, K. et al. Land-use history has a stronger impact on soil microbial community composition than aboveground vegetation and soil properties. Soil Biol. Biochem. 43, 2184-2193 (2011).
View ArticleLi, Q. et al. Biochar mitigates the effect of nitrogen deposition on soil bacterial community composition and enzyme activities in a Torreya grandis orchard. For. Ecol. Manag. 457, 117717 (2020).
View ArticleWaldrop, M. P. & Firestone, M. K. Seasonal dynamics of microbial community composition and function in oak canopy and open grassland soils. Microb. Ecol. 52, 470-479 (2006). PMid:16909344
View Article PubMed/NCBI