Arpita Yadav
Email: arpitayadav@yahoo.co.in
© 2019 Sift Desk Journals. All Rights Reserved
VOLUME: 4 ISSUE: 4
Page No: 476-486
Arpita Yadav
Email: arpitayadav@yahoo.co.in
Sweta Sharma and Arpita Yadav*
Department of Chemistry, University Institute of Engineering and Technology, Chhatrapati Shahu Ji Maharaj University, Kanpur, India 208024
Arpita Yadav, Curcumin and pipernonaline to curb diabetes the natural way: A molecular modeling, docking and dynamics simulation study(2020)Journal of Computational Chemistry & Molecular Modeling 4(4)
This study explores natural compounds derived from Indian spices for their antidiabetic activity in search of safe antidiabetics without side effects. Molecular modeling, docking studies followed by replica exchange molecular dynamics simulation studies have been performed on curcumin, trigonelline, pipernonaline and 6-gingerol to study their PPARγ agonistic activity and α-glucosidase inhibition capability, two prominent targets related to manifestation of diabetes mellitus. It is shown that curcumin and pipernonaline form good, stable complexes with the two targets and hence may be useful as nutraceuticals if adopted alongwith diet and other antidiabetic drugs. They may act as preventive measures against diabetes and are totally safe to consume. They may as well be taken as lead compounds for the further development of better, safe antidiabetics.
Keywords: Curcumin; pipernonaline; natural antidiabetics; PPAR gamma; alpha-glucosidase; Safe antidiabetics
Diabetes: A metabolic disorder
Diabetes mellitus (DM) is one of the most prevalent metabolic disorders around the world. Type 1DM is an autoimmune disease when our body’s immune system destroys the beta cells of pancreas resulting in very little insulin production and hence high blood sugar levels. Type 2 DM which accounts for 80% prevalent cases, occurs when our body fails to respond to normal insulin levels, that is becomes insulin resistant (Berg et al, 2002; Moller, 2001) and hence again results in high blood sugar levels. The long term effects are fatal including myocardial infarction, vascular diseases, stroke etc (www.betterhealth.vic.gov.au). A variety of synthetic drugs are available largely based on targets involved in glucose homeostasis. Clinically used drugs include metformin, sulfonyl ureas, α-glucosidase inhibitors, modulators of sodium glucose cotransporter, thiazolidinediones as PPARγ agonists, GLP-1R agonists and dipeptidyl peptidase 4 inhibitors etc. (Stein et al, 2013). However these synthetic drugs are associated with some side effects (Lee et al, 2019). Rapid increase in number of T2DM patients motivates us to search natural, safe, non-toxic remedies devoid of side effects. Many plant derived natural compounds with antidiabetic property are being used worldwide (Trojan-Rodrigues et al, 2011; Coman et al, 2012). However, natural compound based therapy has not come up as a standalone treatment option. The lack of knowledge about their mode of action and the fact that many natural compounds are multi targeted are some of the barriers in their effective utilization.
In this study we have considered some natural compounds derived from common Indian spices known to possess some antidiabetic activity and tried to evaluate their potential as antidiabetics. It is with the aim that this work may lead to a supplemental, non-toxic, natural antidiabetic to go along with normal diet; we have taken main constituents of fenugreek, black pepper, ginger and turmeric and attempt to understand their Peroxisome proliferator activated receptor (PPARγ) agonistic activity and α-glucosidase inhibition capability. PPARγ is one of the important targets in metabolic disorders to be explored especially if the drug is expected to be multi-targeted. PPARγ regulates adipogenesis and glucose uptake (Zhou et al, 2015). Many flavonoids and similar compounds derived from natural sources are known to target PPARγ (Kuroda et al, 2005) and α-glucosidase (Muraoka et al, 2010; Ieyama et al, 2011; Feng et al, 2011; Chan et al, 2010). However, their mode of action has not been elucidated at the molecular level. An atomistic level understanding would definitely help design a non-toxic, antidiabetic compound from natural leads.
The main constituents of some Indian spices (Niranjan and Prakash, 2008; Grzanna et al, 2005) considered in this study are shown in fig.1. The geometry of these compounds was completely optimized utilizing ab initio quantum mechanical calculations at the Hartree-Fock level with 6-31G basis set (Ditchfield et al, 1971). The optimized conformations are also shown in fig 1. These lowest energy conformations may or may not be the bioactive conformations depending on the conformational flexibility of the compound. The X-ray coordinates of two important targets involved in glucose homeostasis namely PPARγ (pdb file: 4A4V) (De Groot et al, 2013) and α-glucosidase (pdbfile: 5NN8) (Roig-Zamboni et al, 2017) were taken from RCSB protein data bank. The proteins were prepared by adding polar hydrogens using Auto Dock version 4 (Morris et al, 1998) and Auto Dock Vina Version 1.1.2 (Trott and Olson, 2010). Grid for docking ligands was generated and authenticated by redocking the ligand present in X-ray crystal structure with an overall RMSD less than 4Å. Ligands prepared for study were then docked one by one in both the active sites (i.e. PPARγ and α-glucosidase active sites) and their interactions with the active site residues were analysed in detail. AutoDock Vina was used to perform flexible ligand docking studies. Best poses thus obtained were then exported to Shaw’s Desmond software (Bowers et al, 2006) to perform replica exchange simulation studies for better and accelerated sampling of conformational space (Earl and Deem, 2005). Replica exchange molecular dynamics (REMD) simulation studies were performed on best binding energy complexes to judge the dynamical behaviour and stability of complex with time. All atom SPC water was incorporated in a grid of 10x10x10 Å around the protein and orthorhombic boundary conditions were applied. The minimized complex was subjected to simulated annealing followed by dynamics at 300K for 50ns. Parallel trempering swaps between three adjacent replicas were allowed. NPT ensemble was chosen utilizing Martyna-Tobias-Klein barostat method (Martyna et al, 1994). The RESPA integrator was used with 2fs time step for near and bonded and 6fs time step for far atoms (Tuckerman et al, 1992). Coulombic interaction cutoff was set at 9 Å. RMSD analysis has been performed for all heavy atoms together to judge the stability of the system overtime and fluctuations in position of ligand. Deviations have been taken with respect to initial frame for dynamics.
The structures and respective optimized conformations of selected natural compounds are shown in fig. 1. These are largely hydrophobic compounds with some H-bond donor or acceptor groups except trigonelline which occurs in zwitterionic form. After preparing the ligands they were first docked in PPARγ active site. The grid placement in active site is shown in fig. 2. First of all the ligand amorfrutin 2 present in X-ray was redocked to authenticate the grid placement. The best pose interactions of redocked amorfrutin are shown in fig. 3. Docking results for top poses are given in table 1. The ionized carboxylic group of amorfrutin can be seen interacting with arg 288 and cys 285. The hydroxyl group of amorfrutin is also H-bonded to neighbouring polar residues. The hydrophobic chain of amorfrutin interacts with neighbouring hydrophobic residues: gly 284, ile 281, ile341 etc. Fig. 4 shows best poses for interactions of curcumin and 6-gingerol in PPARγ active site. The methoxy groups on curcumin electrostatically interact with arg 288. The –OH groups are also H-bonded to neighbouring hydrophobic residues lining the active site. Detailed contact analysis is presented in table 2.
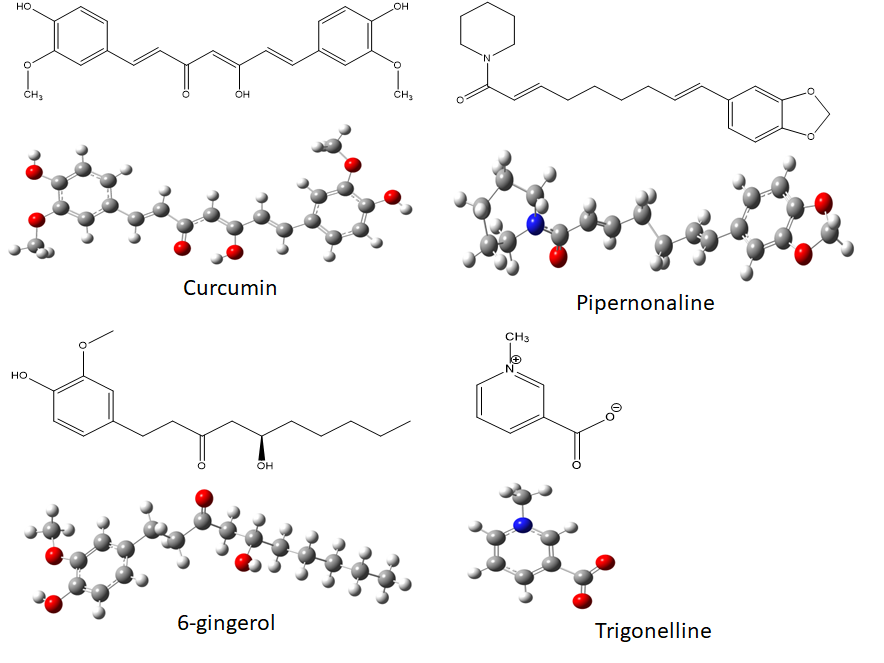
Fig. 1 Chemical structures and optimized conformations of natural antidiabetic compounds
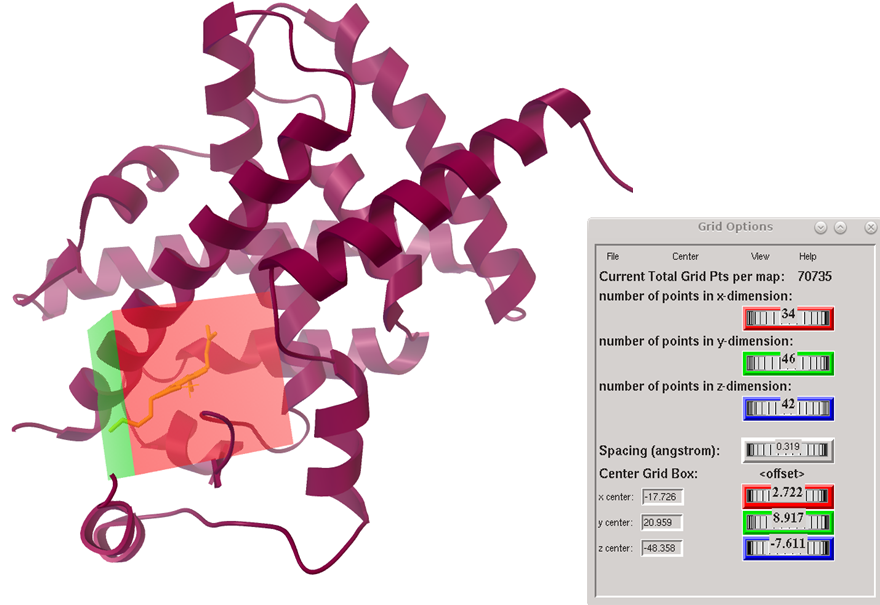
Fig. 2 Grid box placement at the active site of PPAR gamma
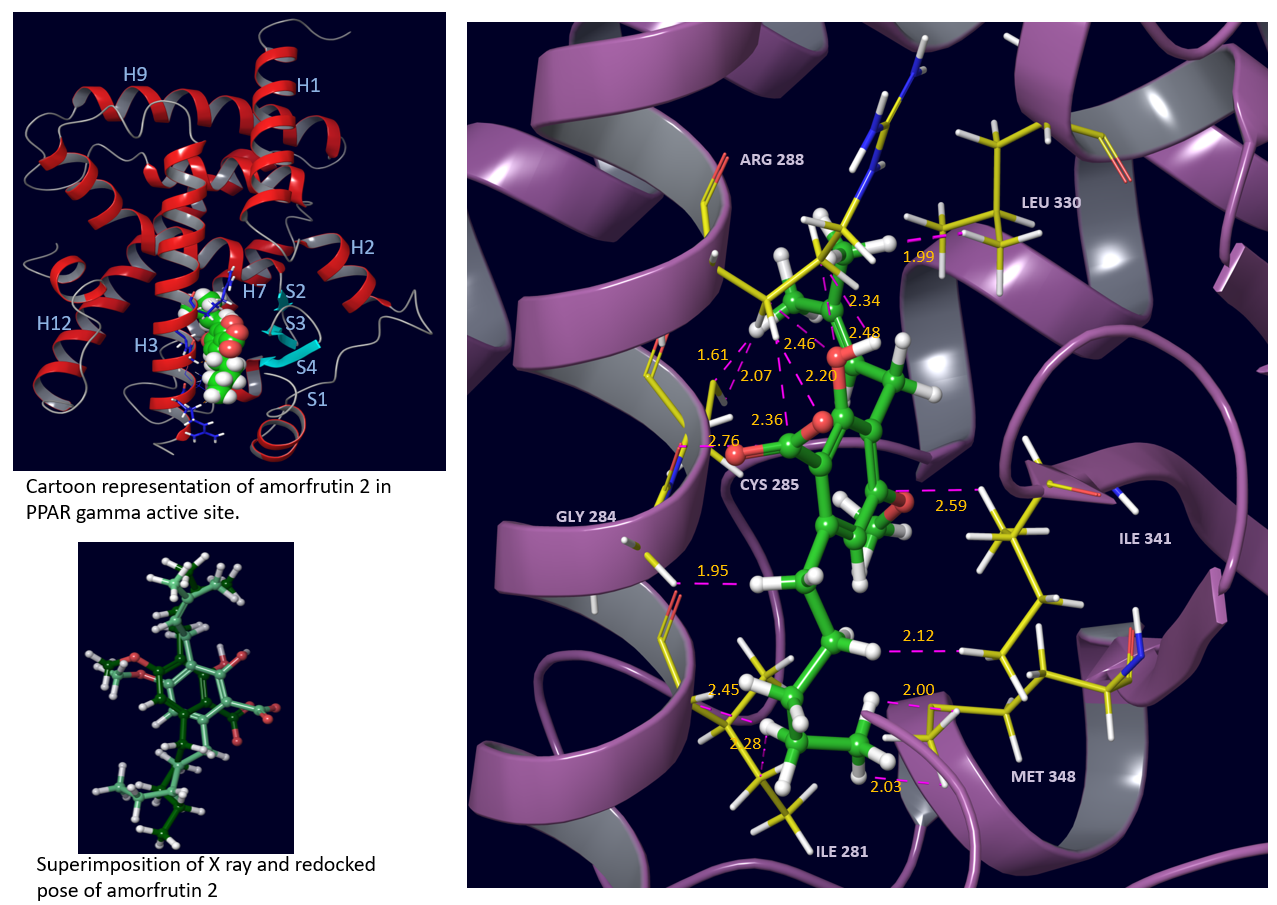
Fig. 3 Closeup view of redocked pose of amorfrutin 2 in PPAR gamma showing interactions with neighbouring residues
|
Table 1. Docking results of natural antidiabetic compounds in PPAR gamma |
|||||||||||||||
|
Redocked amorfrutin 2 |
Curcumin |
6-gingerol |
Pipernonaline |
Trigonelline |
|||||||||||
|
|
Distance from best pose (RMSD in Å) |
|
Distance from best pose (RMSD in Å) |
|
Distance from best pose (RMSD in Å) |
|
Distance from best pose (RMSD in Å) |
|
Distance from best pose (RMSD in Å) |
||||||
|
Pose |
Affinity |
RMSD Lower bound |
RMSD Upper bound |
Affinity |
RMSD Lower bound |
RMSD Upper bound |
Affinity |
RMSD Lower bound |
RMSD Upper bound |
Affinity |
RMSD Lower bound |
RMSD Upper bound |
Affinity |
RMSD Lower bound |
RMSD Upper bound |
|
1 |
-7.4 |
0.000 |
0.000 |
-8.3 |
0.000 |
0.000 |
-7.3 |
0.000 |
0.000 |
-8.2 |
0.000 |
0.000 |
-4.8 |
0.000 |
0.000 |
|
2 |
-7.3 |
3.427 |
8.918 |
-8.1 |
2.207 |
2.938 |
-7.3 |
1.663 |
2.714 |
-7.6 |
2.479 |
9.009 |
-4.8 |
1.245 |
2.181 |
|
3 |
-7.2 |
2.540 |
4.320 |
-8.0 |
7.735 |
10.435 |
-7.3 |
1.051 |
2.016 |
-7.4 |
3.021 |
9.448 |
-4.8 |
3.219 |
4.143 |
|
4 |
-6.9 |
2.786 |
4.962 |
-7.8 |
7.961 |
11.595 |
-7.1 |
1.877 |
2.731 |
-7.1 |
24.125 |
26.029 |
-4.6 |
3.099 |
4.193 |
|
5 |
-6.9 |
3.536 |
5.410 |
-7.7 |
7.853 |
10.724 |
-6.3 |
2.437 |
3.874 |
-7.0 |
18.086 |
22.141 |
-4.6 |
4.918 |
5.666 |
|
6 |
-6.8 |
2.308 |
6.555 |
-7.4 |
2.973 |
3.718 |
-6.3 |
4.571 |
7.239 |
-6.7 |
2.338 |
8.049 |
-4.6 |
4.532 |
5.193 |
|
7 |
-6.8 |
3.256 |
5.779 |
-7.4 |
7.957 |
11.750 |
-6.3 |
3.510 |
7.514 |
-6.7 |
18.789 |
21.908 |
-4.5 |
5.862 |
6.785 |
|
8 |
-6.7 |
2.917 |
8.111 |
-7.3 |
7.576 |
11.412 |
-6.3 |
3.674 |
7.301 |
-6.6 |
24.779 |
26.804 |
-4.4 |
5.592 |
6.919 |
|
9 |
-6.7 |
3.507 |
5.509 |
-7.0 |
6.963 |
11.046 |
-6.2 |
1.739 |
2.426 |
-6.6 |
2.462 |
8.869 |
-4.4 |
14.200 |
15.137 |
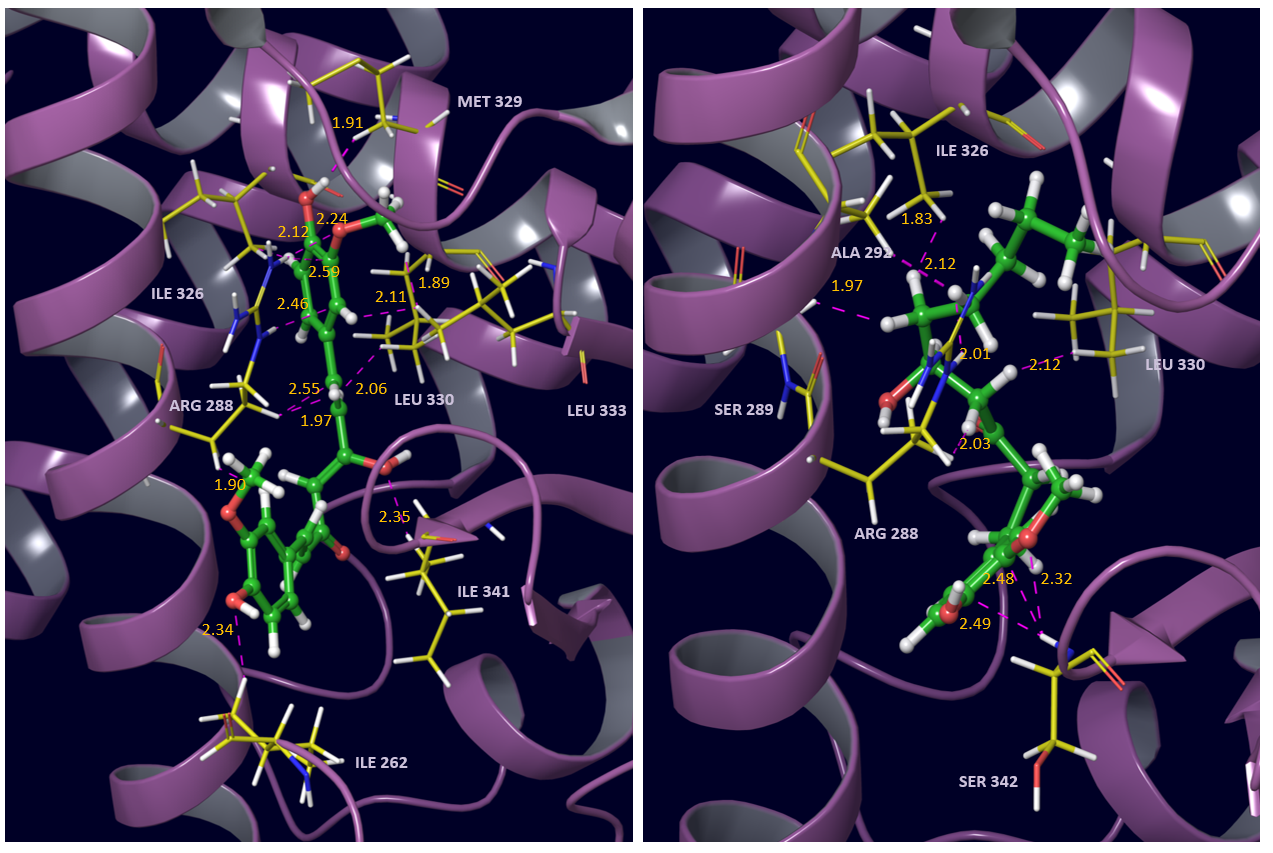
Fig. 4 Interactions of curcumin (left) and 6-gingerol (right) with PPAR gamma active site residues
In 6-gingerol the hydrophobic chain of ligand engages the neighbouring hydrophobic residues of active site which are listed in table 2. The –OH at 6-position interacts with arg 288 though not as efficiently as in curcumin. The substituents on the aromatic ring are H-bonded with the backbone of neighbouring ser 342. Binding affinity observed is comparable to amorfrutin (table1) implying the possibility of activating PPARγ. Docking results for pipernonaline and trigonelline are shown in fig. 5. Trigonelline is held in active site through strong electrostatic interactions with arg 288. Other than electrostatic interactions it is not able to make efficient hydrophobic interactions being very small in size. Its binding affinity (table 1) is also significantly lower than other ligands considered. It is therefore not suited for targeting PPARγ. In pipernonaline the ring oxygens interact electrostatically with arg 288. The hydrophobic chain and the alicyclic ring at the other end interact effectively with the largely hydrophobic pocket of the active site. This is reflected in the binding affinity observed (table 1) comparable to amorfrutin and curcumin. Therefore, out of all the natural anti-diabetics considered curcumin and pipernonaline seem to be the best leads to target PPARγ. It is understood that interactions must be proper for agonism. Considering the similarity of interactions of curcumin/ pipernonaline with PPARγ to amorfrutin with PPARγ; curcumin and pipernonaline are suggested as good, natural, non-toxic lead compounds for the development of safe antidiabetics.
|
Table 2. Detailed contact analysis for best pose of different ligands |
|
|
Ligand |
Interacting residues in PPAR gamma active site |
|
AMORFRUTIN 2 |
ILE281, GLY 284, CYS 285, ARG288, LEU330, ILE341, MET348 |
|
CURCUMIN |
ILE262, ARG288, ILE326, MET329, LEU330, LEU333, ILE341 |
|
PIPERNONALINE |
ILE281, CYS285, ARG288, ALA292, VAL339, ILE341, SER342 |
|
6-GINGEROL |
ARG288, SER289, ALA292, LEU330, ILE326, SER342 |
|
TRIGONELLINE |
ARG288, ILE326 |
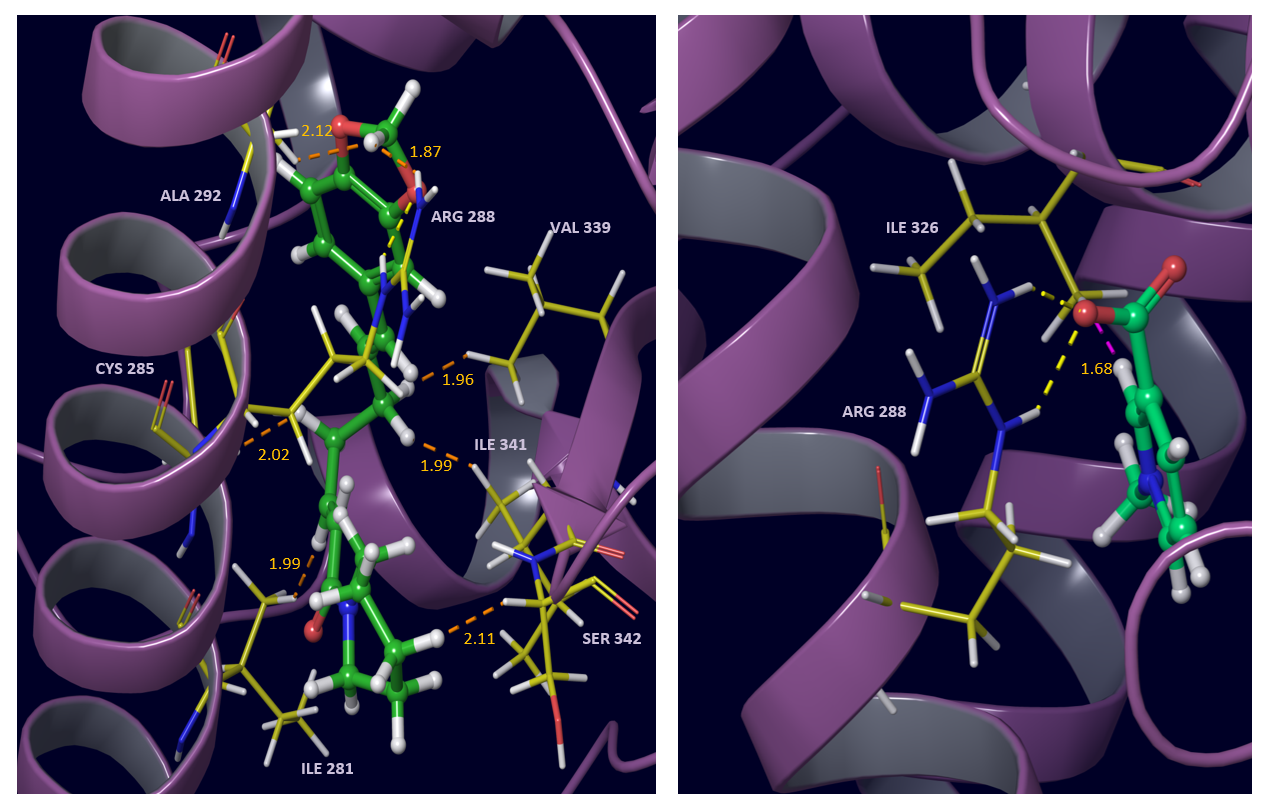
Fig. 5 Interactions of pipernonaline (left) and trigonelline (right) with PPAR gamma active site residues
Apart from PPARγ agonistic activity this study also investigates the α-glucosidase inhibition capability of these natural compounds. As per procedure mentioned in methodology section, the active site of α-glucosidase was prepared. The drug acarbose present in X-ray structure was redocked to authenticate the grid placement. The best pose interactions of redocked acarbose are presented in fig. 6. The results for top poses are given in table 3. The multiple hydroxyl groups of acarbose can be seen H-bonded with neighbouring asp groups of catalytic domain. Complete list of close contacts is given in table 4. In conformity with previous studies (Monique et al, 1991) stating the importance of asp 518 and trp 516; these residues can be seen H-bonded to acarbose. Fig. 7 shows interactions of curcumin and 6-gingerol with active site residues of α-glucosidase. Curcumin interacts similar to acarbose in active site and hence shows similar binding affinity. Instead of asp 518, it utilizes asp 616, trp 481 for anchoring to the active site. The two aromatic rings of curcumin form hydrophobic interactions with neighbouring phenylalanine residues. The binding affinity and interactions indicate the α-glucosidase targeting ability of curcumin. The conformation of 6-gingerol is not appropriate for the active site of α-glucosidase which is evident from its lower binding affinity as well (table 3). 6-gingerol is anchored mainly via H-bonding with trp 613, trp 516 and trp 376. It may target α-glucosidase but not as effectively as curcumin. The docking results for pipernonaline and trigonelline are shown in fig. 8. The benzodioxol ring oxygen of pipernonaline is H-bonded to asp 518. The benzodioxol ring also shows pi-pi stacking with phe 649 and trp 481 aromatic rings. The linker connecting the two aromatic rings in pipernonaline undergoes hydrophobic interactions with neighbouring hydrophobic residues lining the active site. Pipernonaline interacts effectively with α-glucosidase active site residues and shows binding affinity comparable to curcumin and acarbose. It is therefore suited to target α-glucosidase. Trigonelline is a small molecule and does not occupy entire active site but can be seen anchored utilizing H-bonding with trp 516, trp 481 and arg 600. It is encouraging to see that it also blocks the catalytically important trp 481 residue. It implies that it can certainly inhibit α-glucosidase although it may not be highly potent. Again for α-glucosidase inhibition curcumin and pipernonaline seem to be the best natural lead compounds.
|
|
|||||||||||||||
|
Redocked Acarbose |
Curcumin |
6-gingerol |
Pipernonaline |
Trigonelline |
|||||||||||
|
|
Distance from best pose (RMSD in Å) |
|
Distance from best pose (RMSD in Å) |
|
Distance from best pose (RMSD in Å) |
|
Distance from best pose (RMSD in Å) |
|
Distance from best pose (RMSD in Å) |
||||||
|
Pose |
Affinity |
RMSD Lower bound |
RMSD Upper bound |
Affinity |
RMSD Lower bound |
RMSD Upper bound |
Affinity |
RMSD Lower bound |
RMSD Upper bound |
Affinity |
RMSD Lower bound |
RMSD Upper bound |
Affinity |
RMSD Lower bound |
RMSD Upper bound |
|
1 |
-8.0 |
0.000 |
0.000 |
-7.5 |
0.000 |
0.000 |
-5.6 |
0.000 |
0.000 |
-6.7 |
0.000 |
0.000 |
-4.9 |
0.000 |
0.000 |
|
2 |
-7.6 |
3.199 |
5.736 |
-7.4 |
4.865 |
5.646 |
-5.5 |
1.686 |
2.069 |
-6.7 |
4.099 |
6.027 |
-4.9 |
2.653 |
3.767 |
|
3 |
-6.8 |
3.251 |
5.206 |
-7.4 |
3.961 |
10.392 |
-5.5 |
2.514 |
3.392 |
-6.6 |
1.316 |
2.014 |
-4.6 |
1.017 |
1.489 |
|
4 |
-6.5 |
3.714 |
6.076 |
-7.3 |
4.089 |
10.087 |
-5.4 |
2.537 |
6.190 |
-6.6 |
5.732 |
7.936 |
-4.6 |
2.873 |
4.116 |
|
5 |
-6.2 |
3.682 |
10.806 |
-6.7 |
2.615 |
2.812 |
-5.3 |
2.906 |
7.331 |
-6.5 |
2.501 |
8.014 |
-4.3 |
2.764 |
4.413 |
|
6 |
-6.1 |
2.977 |
11.220 |
-6.7 |
1.480 |
10.313 |
-5.3 |
2.492 |
3.379 |
-6.5 |
3.858 |
8.974 |
-4.1 |
2.653 |
4.186 |
|
7 |
-6.0 |
6.128 |
9.293 |
-6.5 |
1.442 |
1.784 |
-5.3 |
2.966 |
4.665 |
-6.5 |
5.005 |
7.750 |
-4.0 |
3.262 |
4.491 |
|
8 |
-5.9 |
3.117 |
10.733 |
-6.4 |
1.846 |
9.869 |
-5.2 |
2.626 |
4.444 |
-6.4 |
1.568 |
1.901 |
-4.0 |
2.942 |
3.946 |
|
9 |
-5.8 |
3.941 |
10.445 |
-6.4 |
6.666 |
8.470 |
-5.2 |
2.640 |
4.327 |
-6.4 |
6.364 |
9.556 |
-3.8 |
7.640 |
8.563 |
|
Table 4. Detailed contact analysis for best pose of different ligands |
|
|
Ligand |
Interacting residues in alpha glucosidase active site |
|
ACARBOSE |
ARG281, ASP282, ASP404, TRP481, TRP516, ASP518, PHE 525, ARG600, ASP616 |
|
CURCUMIN |
ILE441, TRP481, PHE525, TRP613, ASP616, PHE649 |
|
PIPERNONALINE |
ARG411, TRP481, ASP518, PHE649, LEU677, LEU678 |
|
6-GINGEROL |
TRP376, TRP516, TRP613, PHE649, LEU650 |
|
TRIGONELLINE |
TRP481, TRP516, ARG600 |
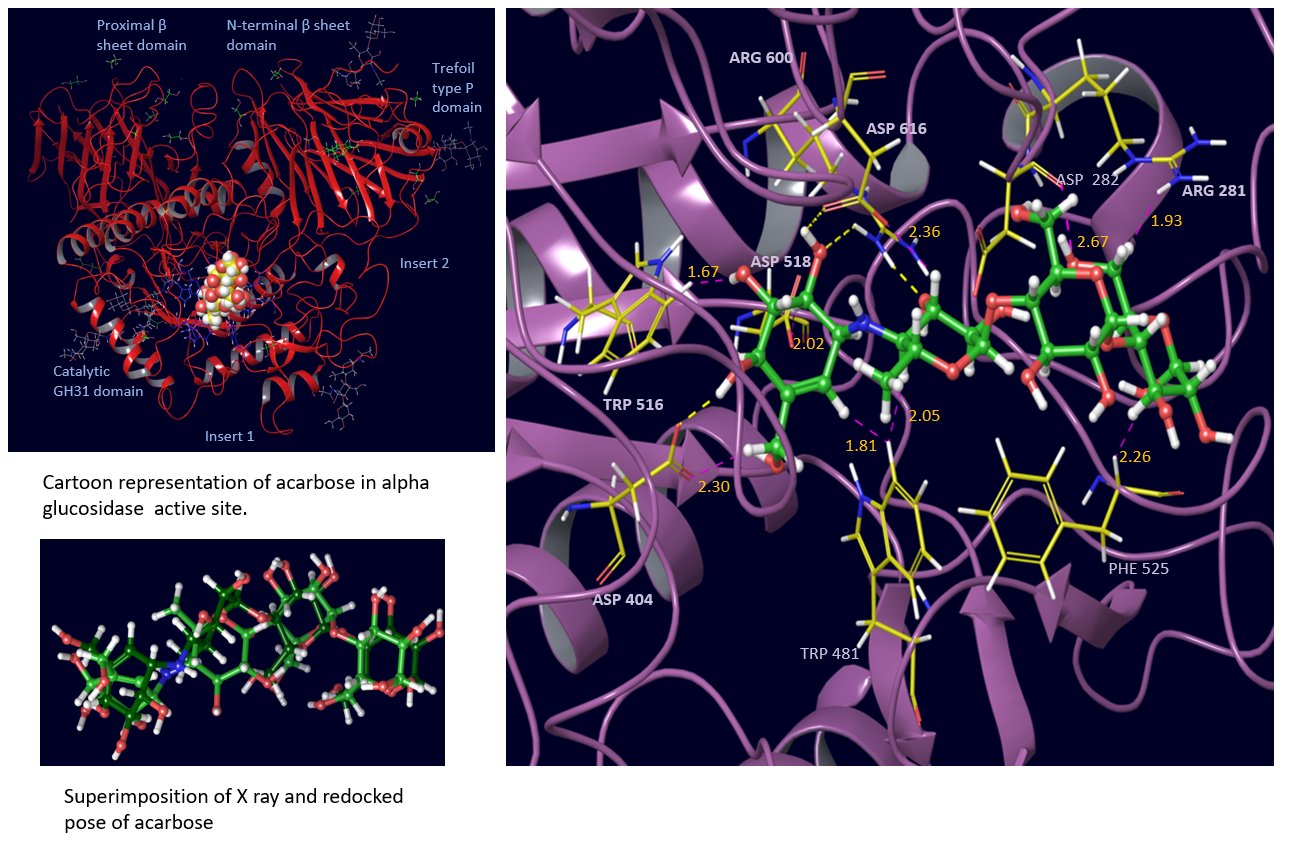
Fig. 6 Closeup view of redocked pose of acarbose in alpha glucosidase showing interactions with neighbouring residues
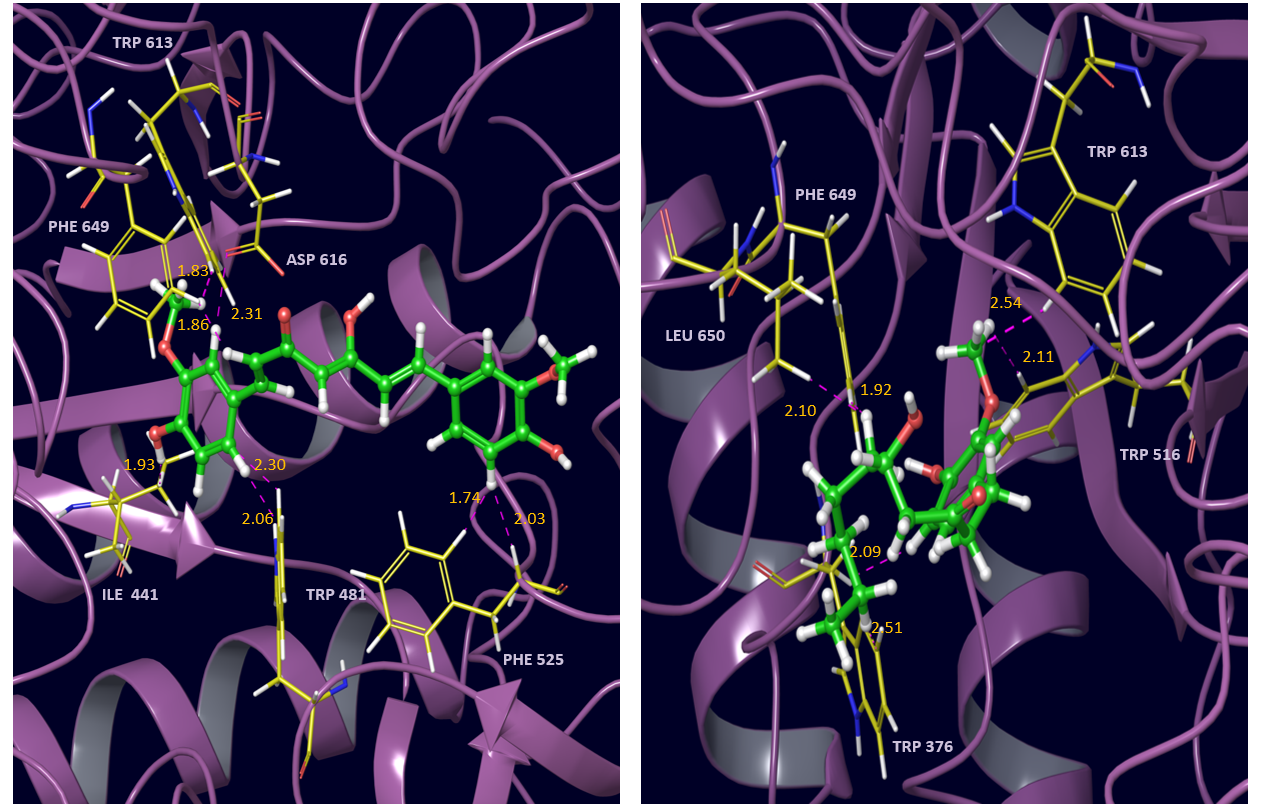
Fig. 7 Interactions of curcumin (left) and 6-gingerol (right) with alpha glucosidase active site residues
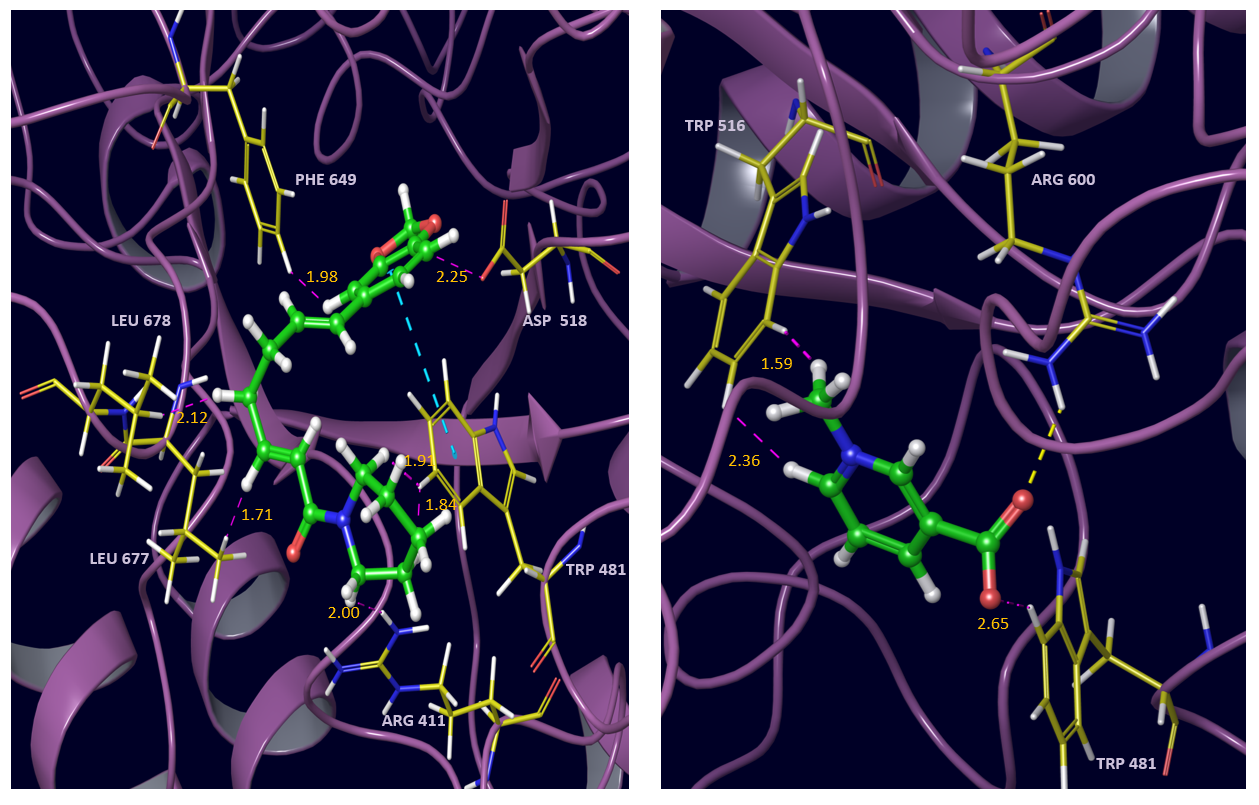
Fig. 8 Interactions of pipernonaline (left) and trigonelline (right) with alpha glucosidase active site residues
We have also investigated the stabilities of best pose complexes of curcumin and pipernonaline in PPARγ and α-glucosidase through REMD simulation studies. Standard MD protocol was followed as mentioned in methodology with three neighbouring replicas. These results are shown in figs. 9 to 11. 50ns trajectories were run in each case. Fig. 9 shows temperature distribution of replicas in temperature spaces for curcumin in PPARγ active site. Fig. 10 indicates good stability of both curcumin and pipernonaline in PPARγ active site. Fig. 11 shows similar results in α-glucosidase. The overall RMSD for all heavy atoms is well below 3Å indicating low fluctuations in position of ligands and good stability of complexes over time for each replica random walk in temperature spaces.
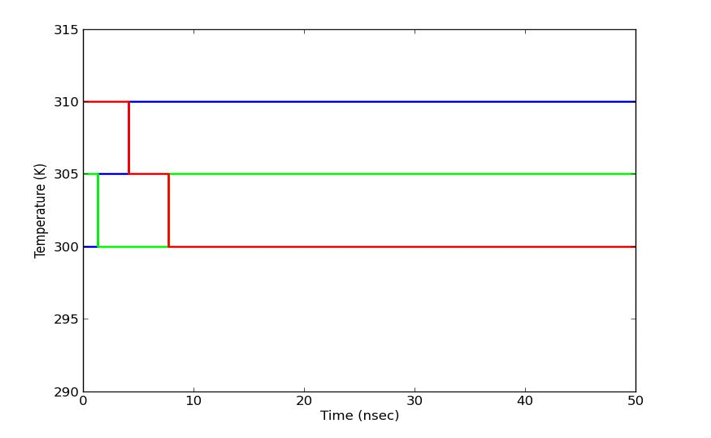
Fig. 9 Temperature distribution for each replica for curcumin in PPAR gamma active site
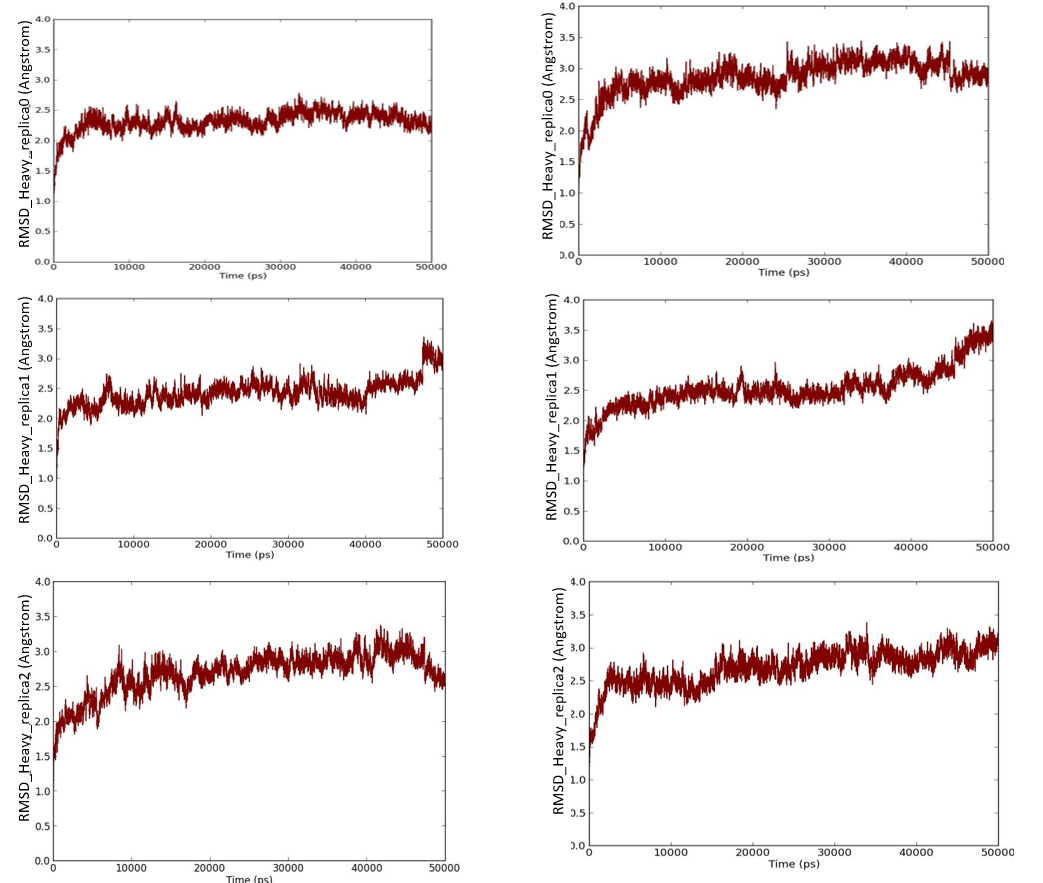
Fig. 10 Root mean square deviation for all heavy atoms with respect to initial frame for best pose complex of curcumin (left) and pipernonaline (right) in PPAR gamma active site
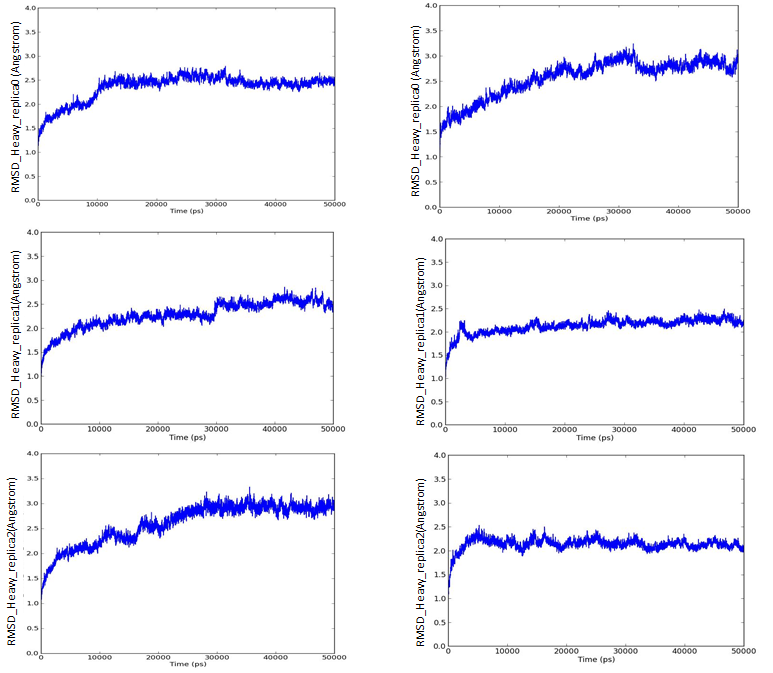
Fig. 11 Root mean square deviation for all heavy atoms with respect to initial frame for best pose complex of curcumin (left) and pipernonaline (right) in alpha glucosidase active site
In this study we have shown at the molecular level how curcumin and pipernonaline can target PPARγ and α-glucosidase similar to other flavonoids and exert antidiabetic action. It is therefore, highly recommended to use curcumin and pipernonaline as nutraceuticals regularly alongwith diet and other antidiabetic drugs to prevent diabetes from worsening and as a preventive measure against its many adverse effects on health.
This study shows at the molecular level the antidiabetic action of two natural, safe compounds curcumin and pipernonaline derived from Indian spices. It is shown that both form stable complexes with PPARγ and α-glucosidase active sites, the two important targets related to diabetes. It is therefore recommended to adopt a nutraceutical made out of curcumin and pipernonaline as a dietary supplement to prevent and overcome the adverse effects of diabetes mellitus type 2. It is also recommended to develop a natural, safe antidiabetic drug from these natural, safe lead compounds.
The authors gratefully acknowledge financial support from Science and Engineering Research Board (SERB), New Delhi (Project no. EMR/2016/000769). Ms. Sweta Sharma also thanks SERB for Senior Research Fellowship. Infrastructural support from University Institute of Engineering and Technology, Chhatrapati Shahu Ji Maharaj University, Kanpur is gratefully acknowledged.
Berg JH, Tymoczko JL, Stryer L (2002) In: Biochemistry, 5th Ed. W.H. Freeman and Company, New York.
Bowers KJ, Chow E, Xu H, Dror RO, Eastwood MP, Gregerson BA, Klepeis JL, Kolossvary L, Moraes MA, Sacerdoti FD, Salmon JK, Shan Y, Shaw DE (2006) Scalable algorithms for molecular dynamics simulations on commodity clusters. Proceedings of the ACM/IEEE conference on supercomputing (SC06) ACM, Tampa, Florida, Nov. 11-17.
View ArticleChan HH, Sun HD, Reddy MVB, Wu TS (2010) Potent α-glucosidase inhibitors from the roots of Panax japonicus CA Meyer var major. Phytochem 71:1360-1364. PMid:20493502
View Article PubMed/NCBIComan C, Rugina OD, Socaciu C (2012) Plants and natural compounds with antidiabetic action. Nat Bot Horti Agrobo 40(1):314-325.
View ArticleDe Groot JC, Weidner C, Krausze J, Kawamoto K, Schroeder FC, Sauer S, Bussow K (2013) Structural characterization of amorfrutins bound to the Peroxisome proliferator-activated receptor gamma. J Med Chem 56: 1535. PMid:23286787
View Article PubMed/NCBIDitchfield R, Hehre WJ, Pople JA (1971) J Chem Phys 54: 724.
View ArticleEarl DJ, Deem MW (2005) Parallel tempering:Theory, applications, and new perspectives Phys Chem Chem Phys 7: 3910-3916. PMid:19810318
View Article PubMed/NCBIFeng J, Yang XW, Wang RF (2011) Bio-assay guided isolation and identification of α-glucosidase inhibitors from the leaves of Aquilaria sinensis. Phytochem 72:242-247. PMid:21215978
View Article PubMed/NCBIGrzanna R, Lindmark L, Frondoza CG (2005) Ginger-an herbal medicinal product with broad anti-inflammatory actions. J Med. Food 8(2):125-132. PMid:16117603
View Article PubMed/NCBIIeyama T, Gunawan-Puteri MDPT, Kawabata J (2011) α-Glucosidase inhibitors from the bulb of Eleutberine americana. Food Chem 128:308-311. PMid:25212136
View Article PubMed/NCBIKuroda M, Mimaki Y, Nishiyama T, Mae T, Kishida H, Tsukawa M, Takahashi K, Kawada T, Nakagawa K, Kitahara M (2005) Hypoglycemic effects of Turmeric (Curcuma longa L. Rhizomes) on genetically diabetic KK-Ay mice. Biol Pharm Bull 28:937-939. PMid:15863912
View Article PubMed/NCBILee JY, Jang SY, Nam CM, Kang ES (2019) Incident hepatocellular carcinoma risk in patients treated with a sulfonylurea: A nationwide, nested case-control study. Scientific reports 9:8532. PMid:31189966
View Article PubMed/NCBIMartyna GJ, Tobias DJ, Klein ML (1994) Constant pressure molecular dynamics algorithms. J Chem Phys 101: 4177-4189.
View ArticleMoller DE (2001) New drug targets for type II diabetes and the metabolic syndrome. Nature 414:821-27. PMid:11742415
View Article PubMed/NCBIMorris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, Olson AJ (1998) Automated docking using a Lamarckian genetic algorithm and on empirical binding free energy function. J Comput Chem 19(14): 1639-1662.
1096-987X(19981115)19:14<1639::AID-JCC10>3.0.CO;2-B
View ArticleMuraoka O, Morikawa T, Miyake S, Akaki J, Ninomiya K, Yoshikawa M (2010) Quantitative determination of potent α-glycosidase inhibitors, salacinol and kotalanol in Salacia species using liquid chromatography-mass spectrometry. J Pharmaceut Biomed 52:770-773. PMid:20303690
View Article PubMed/NCBINiranjan A, Prakash D (2008) Chemical constituents and biological activities of turmeric (Curcuma longa L.). J Food Sci Technol 45(2):109-116.
Roig-Zamboni V, Cobucci-Ponzano B, Iacono R, Ferrara MC, Germany S, Bourne Y, Parenti G, Moracci M, Sulzenbacher G (2017) Structure of human lysosomal acid alpha-glucosidase-a guide for the treatment of Pompe disease. Nat Commun 8: 1111. PMid:29061980
View Article PubMed/NCBIStein SA, Lamos EM, Davis SN (2013) A review of the efficacy and safety of oral anti-diabetic drugs. Expert Opin Drug Safety 12:153-175. PMid:23241069
View Article PubMed/NCBITrojan-Rodrigues M, Alves TLS, Soares GLG, Ritter MR (2011) Plants used as antidiabetics in popular medicine in Rio Grande do Sul, Southern Brazil. J Ethnopharmacol 139(1):155-163. PMid:22079795
View Article PubMed/NCBITrott O, Olson AJ (2010) AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J Comput Chem 31(12): 455-461. PMid:19499576
View Article PubMed/NCBITuckerman M, Berne BJ, Martyna GJ (1992) Reversible multiple time scale molecular dynamics. J Chem Phys 97(3): 1990-2001.
View ArticleZhou T, Yan X, Wang G, Liu H, Gan X, Zhang T, Wang J, Li L (2015) Evolutionary pattern and regulation analysis to support why diversity functions existed within PPAR gene family members. Bio Med Res Int 11 Article ID 613910. PMid:25961030
View Article PubMed/NCBIMonique H, Kroos M, Beeuman JV, Oostra B, Reuser A (1991) Human lysosomal α-glucosidase characterization of the catalytic site. J Biol Chem 21: 266(21): 13507-13512.
