Mr. Sajjad Esmaeili
E-mail: sajad.smaeili@kums.ac.ir
© 2019 Sift Desk Journals. All Rights Reserved
VOLUME: 4 ISSUE: 2
Page No: 392-402
Mr. Sajjad Esmaeili
E-mail: sajad.smaeili@kums.ac.ir
Yazdan Rahmatia, Hasan Mollanooria, Soodabeh Shafieeb, Sajjad Esmaeilic*
aDepartment of Medical Genetics, Iran University of Medical Sciences, Tehran, Iran
bDepartment of Biology, Sciences & Research Branch, Islamic Azad University, Tehran, Iran
c Medical Biology Research Center, Health Technology Institute, Kermanshah University of Medical Sciences, Kermanshah, Iran
Yazdan Rahmati, Soodabeh Shafiee, Sajjad Esmaeili, Identification of 7 key age-related genes involved in Kawasaki disease, an integrated study by metaDE and weighted gene co-expression network analysis(2020)Journal of Computational Chemistry & Molecular Modeling 4(2) pp:392-402
Kawasaki disease (KD) is an inflammatory vasculitis of unknown etiology that occurs in infantile and shows a higher incidence in younger children. This work aimed to investigate the correlation between gene expression status and age of disease onset by integrated bioinformatics analysis. In the first step by using metaDE package, meta-analysis performed for three expression arrays including GSE73464, GSE18606, and GSE68004. In the next step, WGCNA package applied on meta-analysis results for the detection of genes in modules that correlate with the gender and the disease age of onset. Finally, Functional annotations of these genes were carried out to highlight the KD-associated molecular pathways. Our meta-analysis identifies 2417 differentially expressed genes as the most important genes. By applying WGCNA upon these genes we identified a module that has a negative correlation with disease age of onset, so genes in this module show an important role in KD. Functional annotations revealed inflammatory pathway and immune response pathways to infections as the most enriched terms. Genes including, CHUK, TLR5, TICAM2, LY96, MYD88, IRAK4, and JAK2 identified as hub genes. Among them the role of LY96, TLR5, MYD88, and IRAK4 in KD had been confirmed in previous studies but for the first time we introduce TICAM-2, CHUK, and JAK2 as genes that can have important role in KD.
Keywords: Kawasaki disease, metaDE, WGCNA, gene expression, age of onset
KD is an inflammatory vasculitis of unknown etiology that occurs in infantile and toddlers. Its main complication is the development of coronary artery lesions (CALs) including coronary arterial dilation, stenosis, and aneurysms [1-3]. Kawasaki disease (KD) exhibits many signs such as high temperature, erythema which occurs in the mucosa of lips or mouth, alterations in the body organs, and large lymph nodes in the neck [4-6]. KD among Japanese and Japanese American children is clearly more common (265/100,000) than other populations like other Asian countries (51 to 194/100,000), Europe (8.39/100,000) or American children (20.8/100,000) [7-12]. To date, the etiology of KD has remained an undetermined question and the causative agents are also unexplained [13]. But, based on researchers’ findings, a genetic predisposition considered as a cause to the development of KD and also the interaction of an unknown infectious source can predispose to this disease [14]. The observation of familial occurrence and raised frequency in Asian populations, taken together suggest the existence of a genetic basis [15, 16]. Associations with genetic variation in several genes such as BLK, CASP3, CD40, FCGR2A, IPTKC, and HLA class II have been ascertained in diverse populations. Also, it is known that the increased risk for the development of coronary aneurysms in populations with European ethnicity is correlated with the genetic variation in the TGF pathway (TGFβ2, TGFβR2, SMAD3) [17].
Heart damage is visible in a significant number of untreated KD patients and they can develop myocardial infarction, unexpected death, or ischemic heart disorder from ectasia (Coronary artery aneurysms) [18, 19]. Early diagnosis is crucial for effective treatment which results in abolition of the inflammatory process to reduce the risk of coronary artery aneurysms (CAA) rates to approximately 5% to 10% [20]. CAA is the most important complication of KD patients and untreated KD children, the disease-associated inflammation modifies the arterial wall which leads to CAA in 25% of them [21]. In developed countries, KD has been reported as the most prevalent cause of acquired heart disease in children [22]. KD shares certain features to other childhood febrile conditions, including, infectious (e.g. staphylococcal and streptococcal toxic shock syndromes, measles and other viral illnesses) and inflammatory conditions which makes differential diagnosis difficult [20]. Although nowadays there are guidelines to assist diagnosis based on clinical signs and symptoms, such as echocardiography, and laboratory variables, rapid diagnosis of KD from other mimicking conditions for treatment and impediment of CAA development remains a momentous mission [23].
Recently, analysis algorithms for the meta-analysis of array data and differential co-expression network, advanced and implemented to study the expression data of genes and microRNAs [24-26]. There is a new functional strategy in systems biology, the weighted gene co-expression network analysis (WGCNA) algorithm, which identifies the most important genes in co-expressed genes that are associated with a sample trait [27, 28]. WGCNA makes modules, sub-network regions, based on similarities in expression profiles of samples, and detects associated genes [29, 30]. By analysis of these modules in two different conditions, healthy control against disease samples, we aimed to identify gender and age of onset -related genes that may represent potential diagnostic biomarkers as well as therapeutic targets with clinical utility.
2.1. Expression array datasets and primary processing
Three expression datasets including GSE73464, GSE18606, and GSE68004 in total studied for gene expression of 86 controls 194 patients. These data sets obtained from GEO NCBI at https://www.ncbi.nlm.nih.gov/geo by searching Kawasaki disease and Homo sapiens in the query line. GSE73464 and GSE68004 are produced using Illumina HumanHT-12 V4.0 expression beadchip and included 77 healthy and 154 disease samples. To normalize these datasets, quantile normalization method in limma package was used. GSE18606 produced using Agilent-014850 Whole Human Genome Microarray 4x44K G4112F and included 9 healthy and 40 disease samples. To normalize this dataset quantile normalization method in limma package was used too. For better results, we included only single measure of each gene by the aggregate function in the S4Vectors package, which gives an average measure of the probes of each gene.
2.2. Quality control, comparability and detection of outlier samples
By metaQC [31] package quality of datasets are assessed and samples that have low quality removed from the rest of analysis, p-value cutoff 0.05 was used for the selection of differentially expressed genes, and p-value cutoff 0.05 was used for the selection of pathways by metaQC package. To evaluate the comparability of control and disease samples, soft Connectivity function in WGCNA package was used. Soft Connectivity evaluates two factors, 1) correlation of expression level of each gene between two datasets, 2) correlation of connectivity of each gene between two datasets. The datasets are comparable if two mentioned correlations are positive and have significance p.value. To remove outlier samples, standardized connectivity (Z. K) method used and samples which had Z. K score< −2 deleted from the rest of the analysis.
2.3. Meta-analysis and detection of significant genes
Fisher method in metaDE package was used to recognize significant genes. FDR cutoff 0.001, used to access the most important genes in this study.
2.4. Network construction and module detection by WGCNA package
The results of metaDE set as input for WGCNA package. The disease dataset of GSE73464 used as the main dataset for the rest of analysis. One weighted gene co-expression networks according to disease samples was made. Using pickSoftThreshold function that helps to choose proper soft-thresholding power, we selected soft thresholding power of 6 for providing scale-free topology fit index that reaches values above 0.9. After calculation of adjacencies, to minimize the effect of noise and spurious associations, adjacency results transformed to Topological Overlap Matrix (TOM), then scaling of Topological Overlap Matrices was used to mitigate the effect of different statistical properties. We used Quantile-quantile plot of the TOMs in the dataset to see what scaling is achieved. Results of TOM set as input to produce dendrogram of genes. CutreeDynamic function used for branch cutting. Minimum module size of 30, and the module detection sensitivity deep Split 2 in blockwise Consensus Modules function were used for network construction.
2.5. Identification of clinically significant modules and functional annotation
Module membership (MM) and gene significance (GS) for evaluating correlation between genes and traits were measured. GS is the correlation between genes and traits and MM is the correlation of the module eigengene and gene expression profile. By measuring MM and GS, we can identify modules with high module membership as well as significant genes related to gender and age of onset. We also evaluated the correlation between GS and MM for genes in modules with high module membership in which central genes considered as the most important elements of the module associated with the trait.
2.6. Detection of hub genes and their functional annotations
Within detected module, hub genes were screened according to criteria cor.geneModuleMembership > 0.8 and cor.geneTraitSignificance > 0.25. Next we carried out functional enrichment analyses of known genes in order to facilitate the interpretation of the biological mechanisms related to these genes. KEGG and STRING databases used to functional and biological interpretation of detected genes, using the KEGG database, important signaling pathways with P.value < 0.05 and combined score > 10 were identified. PPI network of detected genes was constructed by Search Tool for the Retrieval of Interacting Gene (STRING10.5; https://string-db.org/) with a combined score >0.4 as the cut-off point. The coexpression network of the genes was constructed by Cytoscape software[32].
3.1. Quality control, comparability and detection of outlier samples
The quality of the datasets was examined using MetaQC package. According to the results of the metaQC package and especially SMR score, we decided to keep all selected datasets in the process of analysis (Table. 1). Dataset comparability evaluated by the soft Connectivity function that provides positive correlation values when there is a high comparability between control and disease datasets for each expression array. Our datasets were comparable as the overall gene expression correlation for GSE73464 was (cor=0.99, p<1e−200), the overall gene expression correlation for GSE68004 was (cor=0.98, p<1e−200) and for GSE18606 was (cor=0.72, p<1e−200) (Fig.1). After selecting GSE73464 as the final dataset for WGCNA package analysis, based on standardized connectivity (Z. K) method, 3 outlier samples removed from this dataset. Figure 3A shows the correlation between clinical traits and the sample dendrogram depicted for the disease dataset of GSE73464.
Table 1: quality control of three datasets using metaQC package. . IQC, internal quality control; EQC, external quality control; CQCg, consistency quality control of gene; CQCp, consistency quality control of pathway; AQCg, accuracy quality control of gene; AQCp, accuracy quality control of pathway, SMR, standardized mean rank.
|
datasets |
Groups
|
IQC |
EQC |
AQCg |
AQCp |
CQCg |
CQCp |
SMR |
|
GSE73464 |
Control-patient |
5.615 |
6.52 |
248.5 |
5.622 |
568.6 |
410 |
1.5 |
|
GSE68004 |
Control-patient |
3.3 |
3.594 |
204.5 |
24.77 |
454.4 |
410 |
2.25 |
|
GSE18606 |
Control-patient |
5.615 |
5.661 |
68.24 |
1.161 |
94.86 |
410 |
2.5 |
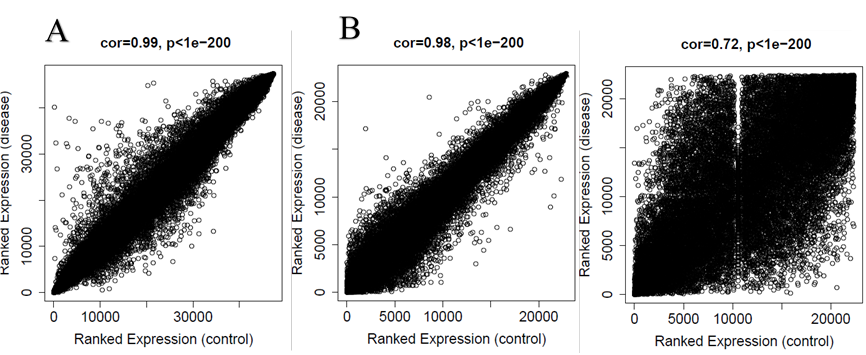
Figure 1: correlations and the p-values between control and disease datasets for (A) GSE73464, (B) GSE68004 and (C) GSE18606. As it is clear correlations and the p-values for all datasets are positive and significance.
3.2. Meta-analysis and detection of significant genes
The results of MetaDE package, a heatmap that present up and down regulated genes is summarized in Figure 2. 2417 DEGs using FDR cutoff 0.001 detected from which 930 are upregulated and 1487 are downregulated. These genes set as input for WGCNA package.
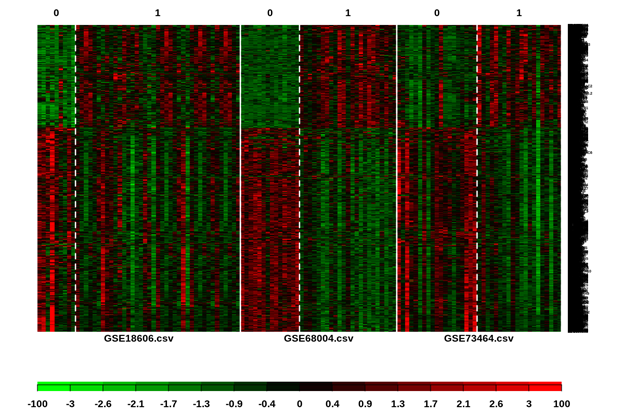
Figure 2: Hierarchical clustering heat map of samples from control-patients. The gene expression heat map of the 2417 differentially expressed genes for the control-patient groups. Red and green indicate high and low expression in samples.
3.3. Network construction and module detection by WGCNA package
The KD dataset related to GSE73464 exhibited a scale-free topology as the power was set to 6 and the Scale-free Topology Fit Index reached values above 0.9 for low powers (< 30). . This also shows that the batch-effects was not present in our dataset (Fig.4). For module identification, weighted gene co-expression networks constructed for this dataset. Based on the WGCNA package, a module is a group of strongly co-expressed genes, these genes have similar biochemical and functional properties or belong to similar pathways. By hierarchical clustering, 8 modules identified for the KD dataset, these modules have different sizes in terms of the number of genes which are labelled by the different colors and shown in figure 3B.
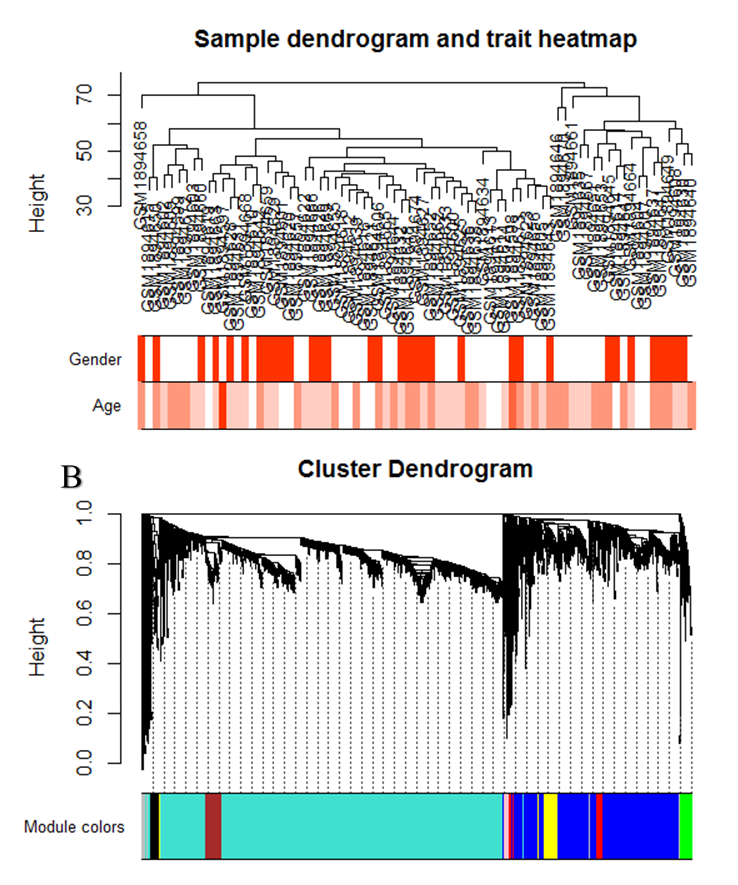
Figure 3: (A) Clustering dendrogram of samples based on their Euclidean distance for GSE73464 disease dataset. (B) Clustering dendrogram of genes, with dissimilarity based on topological overlap, together with assigned module colors for disease dataset.
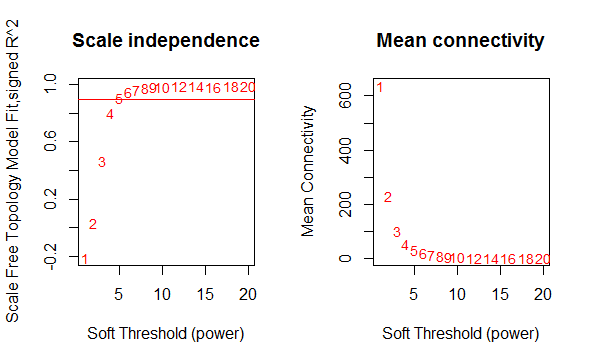
Figure 4: Analysis of network topology for various soft-thresholding powers.
3.4. Identification of clinically significant modules and functional annotation
To identify modules that are significantly associated with the measured clinical traits (Age of onset, Gender), we used Module-trait association plot. The analysis identifies several significant module–trait associations. As it can be seen from the plot, the best correlation exists between age of onset and the yellow module (Fig. 5A). We quantified association between genes of the yellow module and age of onset by measuring GS and MM (Fig. 5B). A heatmap was made to analyze the interaction between 8 modules, the heatmap can depict adjacencies or topological overlaps, with light colors denoting low adjacency (overlap) and darker colors higher adjacency (overlap) (Fig. 5C). we used plot Eigengene Networks function in WGCNA package to generates a summary plot of the eigengene network and also add a clinical trait (Age of onset ) to the eigengenes to see how the traits fit into the eigengene network (Fig. 5D).
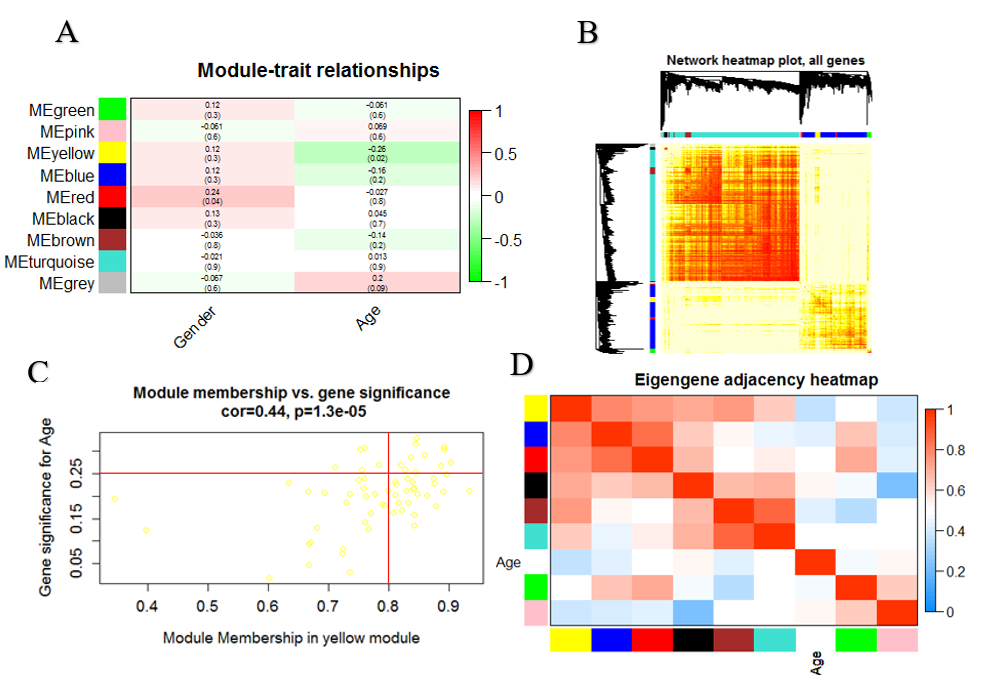
Figure 5: Identification of modules associated with the clinical traits: (A) Heatmap of the correlation between module eigengenes and clinical data. The yellow module was significantly correlated with age. (B) Scatter plot of module eigengenes in yellow module. (C) Interaction relationship analysis of co-expression genes. Different colors of horizontal axis and vertical axis represent different modules. The brightness of yellow in the middle represents the degree of connectivity of different modules. (D) Module eigengene dendrogram and eigengene network heatmap summarize the modules yielded in the clustering analysis.
3.5. Identification of hub genes and functional analysis
We find yellow module as the most important one in correlation of KD and age of onset, genes related to the yellow module can be seen in Table 2. According to the results of KEGG database analysis, the most important signaling pathways related to the genes of yellow module are shown in Figure 6A, the cenplot that shows the relation between signaling pathways and yellow module genes are shown in Figure 6B. We set out to identify hub genes with the highest module membership scores according to the criteria cor.geneModuleMembership > 0.8 and cor.geneTraitSignificance > 0.25 in yellow module, then to identify the most significant clusters of the hub genes, PPI network was constituted by STRING as shown in Figure 7. Genes like CHUK, TLR5, TICAM2, LY96, MYD88, IRAK4, and JAK2 are the most interacted genes which are identified as hub genes.
Table 2: the list of genes identified in yellow module related to KD
|
Module |
genes |
|
Yellow |
SCO2, WAC, ZFP36L1, KDM3B, SPG11, PJA2, SYNJ1, ZNF281, PARP9, NT5C2, UBQLN2, RFWD2, SUMO1P3, WDR51B, RAB33B, FEM1C, KBTBD7, MTMR6, TMEM71, FAM45A, CYB5R4, TLR5, HMGCR, WDR26, JAK2 , GOLPH3, TP53INP1, STXBP5, SLK, IRAK4, TMEM188, NIN, SCYL2, PAPD4, USP15, TLE4, CCDC128, AADACL4, MIER1, FBXO30, CHUK, FCHO2, SMNDC1, USP6, SEL1L, PTPN12, CMTM6, OSBPL11, PCMT1, RHOT1, LY96 , VCPIP1, ZFYVE16, MTHFD2, RP5-1022P6.2, BAZ2B, DCP2, LYPLA1, SLC16A6, RAB11A, ZMPSTE24, IPO11, ZEB2, CNIH4, IFRD1, TICAM2, MyD88 |
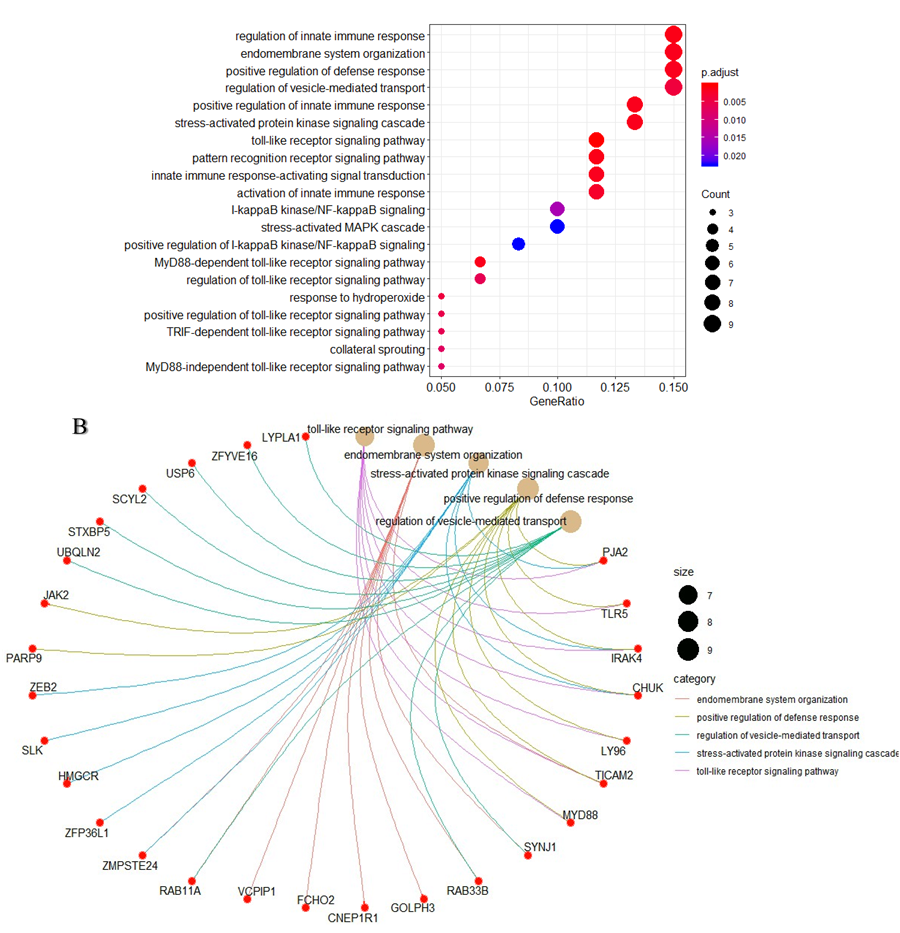
Figure 6: Functional enrichment analysis of yellow module: (A) KEGG pathway analysis of all genes in yellow module. (B) cnetplot of all genes in yellow module that depicts the linkages of genes and most important signaling pathways.
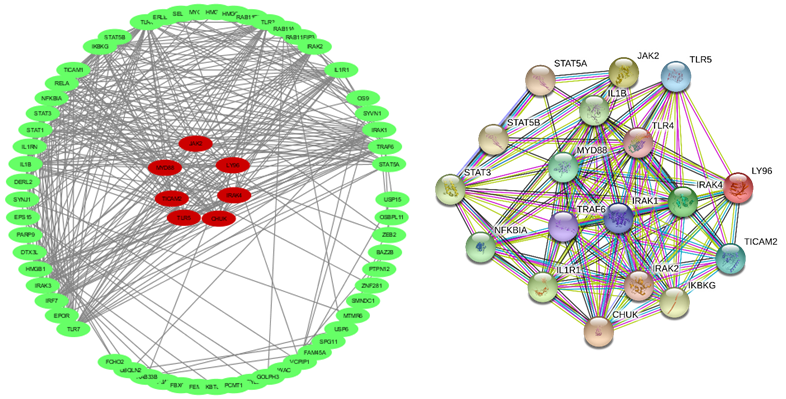
Figure 7: The PPI network of detected hub genes that was analyzed by String software. Genes like CHUK, TLR5, TICAM2, LY96, MYD88, IRAK4, and JAK2 have important role in this network.
KD is one of the rare diseases that can become very serious and deadly due to the involvement of human inflammatory pathways and blood vessels [1, 18]. It is important to note that KD is seen only in people of a low age and its prevalence in Asia is higher than in other parts of the world. According to studies in the field, inflammatory pathways are activated in innate immune cells. Studies have shown that inflammatory pathways are activated in innate immune cells and stimulate pro-inflammatory cytokines such as TNF-β. In this regard, there have been studies that target these signaling pathways and genes that have influenced the course of the disease. However, the disease is still considered pathologically unknown [1, 33, 34]. In this study by selecting three datasets that all of them evaluate the difference of expression between control and disease samples, the correlation between DEGs and Age of onset of KD explored. At first step, it tried to identify the most important genes by metaDE package. We identified 2417 DEGs in two up and down regulated groups. Because of high quality and much number of disease samples, GSE73464 dataset selected for the rest of the analysis, so expression status of DEGs in disease samples of the mentioned dataset used as input for WGCNA package, we also considered age of onset and gender of disease samples as clinical data in our analysis. As seen in figure 3-A, the most valuable correlation between clinical traits and detected modules is a negative association between age of onset and yellow module. It seems genes in yellow module play an important role in disease with early age of onset. We constructed modules and subgrouped regions based on similarity in expression profiles of samples and gene links using WGCNA method. These modules are designed to identify the strong relationships with the onset and development of KD so that they can be used disease treatment. According to Figure 3, the yellow modulus was considered as a module with high association with the disease. All 67 yellow module genes are listed in Table 2. The genes with the highest module membership scores and also with the most number of interactions with other genes related module considered hub genes of that module. Enrichment was then carried out on the basis of signaling pathways (by KEGG pathways; https://www.genome.jp/kegg/pathway.html) and protein-protein interactions (PPI) by STRING software.
Finally, 7 of these genes including CHUK, JAK2, TLR5, TICAM2, LY96, MYD88 and IRAK4 were selected because of the high score. MYD88 (Myeloid differentiation primary response 88) is involved in the signaling pathway within the immune cells as an adapter, it activates transcription factor NF-κB by TLRs and IRAKs [35, 36]. One study found that the activation of MyD88-dependent inflammatory signals in macrophages and vascular cells led to a negative effects on KD [37]. MYD88 also activates pro-inflammatory cytokines such as TNF-α by activating the TLR4-MYD88 signaling pathway [38]. TLR5 (Toll-like receptor 5) have been found to be involved in the onset of many diseases, such as inflammatory bowel disease [39],. TLR5 is present in the cell membrane and through MYD88 and IRAK4, it activates the innate immune system and secretes cytokines such as TNF-α [40-42]. One study showed that the Toll-like receptor family, including TLR5, was upregulated in KD [43].
LY96 (lymphocyte antigen 96) as called myeloid differential protein-2 (MD-2) along with TLR5 is involved in the regulation of TNF-α and nuclear factor-κB signaling pathway [44, 45].
JAK2 (Janus kinase-2) is a non-receptor tyrosine kinase involved in cytokine receptors signaling pathways such as interferon receptors, and GM-CSF receptor family. Past studies have shown that JAK2 has been implicated in many diseases, such as leukemia [46] and myelofibrosis, but has not been reported in KD [47]. CHUK (Conserved helix-loop-helix ubiquitous kinase), also called IKKα, is a nuclear factor-κB transcription activator, and it leads to stimulates lipogenesis in the context of hepatitis C virus infection [48]. Also it is involved in signaling pathways of the MAPK, adipocytokine, PI3K-Akt, and mTOR associated with lipid metabolism, but has not been reported in KD [49].
TICAM-2 (TIR-containing Adapter Molecule), also called MYD88-4 or TIRP involved in toll receptor signaling. One study concluded that TICAM-2 is involved in the process of INF-B production through TICAM-1 [50].
Our results shows that LY96, TLR5, MYD88, and IRAK4 genes are important in KD etiology. On the other hand, TICAM-2, CHUK, and JAK2 genes are the first to be reported in relation to KD. According to the purpose of the study, these genes are predicted to play a critical role in the onset and development of KD and can be potential target in the therapeutic process. Finally, this study should be substantiated by evaluating the expression levels of these genes and proteins in clinical samples.
In this study for the first time, correlation of several genes with Kawasaki disease in earlier ages detected, these genes are TICAM-2, CHUK, LY96, TLR5, MYD88, IRAK4 and JAK2. This information helps understand the molecular mechanism and treatment of KD.
Burns, J.C. and M.P. Glodé, Kawasaki syndrome. The Lancet, 2004. 364(9433): p. 533-544. 16814-1
View ArticleKuo, H.C., Preventing coronary artery lesions in Kawasaki disease. Biomed J, 2017. 40(3): p. 141-146.
View ArticleMori, M., et al., Two-generation Kawasaki disease: mother and daughter. The Journal of pediatrics, 2001. 139(5): p. 754-756. PMid:11713463
View Article PubMed/NCBINewburger, J.W., et al., Diagnosis, treatment, and long-term management of Kawasaki disease: a statement for health professionals from the Committee on Rheumatic Fever, Endocarditis and Kawasaki Disease, Council on Cardiovascular Disease in the Young, American Heart Association. Circulation, 2004. 110(17): p. 2747-2771. PMid:15505111
View Article PubMed/NCBIKawasaki, T., Acute febrile mucocutaneous syndrome with lymphoid involvement with specific desquamation of the fingers and toes in children. Arerugi, 1967. 16: p. 178-222.
Kato, H., et al., Long-term consequences of Kawasaki disease: a 10-to 21-year follow-up study of 594 patients. Circulation, 1996. 94(6): p. 1379-1385. PMid:8822996
View Article PubMed/NCBIMakino, N., et al., Descriptive epidemiology of Kawasaki disease in Japan, 2011-2012: from the results of the 22nd nationwide survey. J Epidemiol, 2015. 25(3): p. 239-45. PMid:25716368
View Article PubMed/NCBILue, H.C., et al., Estimation of the incidence of Kawasaki disease in Taiwan. A comparison of two data sources: nationwide hospital survey and national health insurance claims. Pediatr Neonatol, 2014. 55(2): p. 97-100. PMid:23890670
View Article PubMed/NCBIDu, Z.D., et al., Epidemiologic study on Kawasaki disease in Beijing from 2000 through 2004. Pediatr Infect Dis J, 2007. 26(5): p. 449-51. PMid:17468660
View Article PubMed/NCBIKim, G.B., et al., Epidemiology and Clinical Features of Kawasaki Disease in South Korea, 2012-2014. Pediatr Infect Dis J, 2017. 36(5): p. 482-485. PMid:27997519
View Article PubMed/NCBIHarnden, A., et al., Kawasaki disease in England: ethnicity, deprivation, and respiratory pathogens. Pediatr Infect Dis J, 2009. 28(1): p. 21-4. PMid:19145710
View Article PubMed/NCBIHolman, R.C., et al., Kawasaki syndrome hospitalizations among children in Hawaii and Connecticut. Archives of pediatrics & adolescent medicine, 2000. 154(8): p. 804-808. PMid:10922277
View Article PubMed/NCBIShulman, S.T. and A.H. Rowley, Kawasaki disease: insights into pathogenesis and approaches to treatment. Nat Rev Rheumatol, 2015. 11(8): p. 475-82. PMid:25907703
View Article PubMed/NCBIRamphul, K. and S.G. Mejias, Kawasaki disease: a comprehensive review. Archives of medical sciences. Atherosclerotic diseases, 2018. 3: p. e41-e45. PMid:30775588
View Article PubMed/NCBINakamura, Y., et al., Epidemiologic features of Kawasaki disease in Japan: results of the 2009-2010 nationwide survey. J Epidemiol, 2012. 22(3): p. 216-21. PMid:22447211
View Article PubMed/NCBIDergun, M., et al., Familial occurrence of Kawasaki syndrome in North America. Arch Pediatr Adolesc Med, 2005. 159(9): p. 876-81. PMid:16143748
View Article PubMed/NCBIHedrich, C.M., A. Schnabel, and T. Hospach, Kawasaki Disease. Frontiers in pediatrics, 2018. 6: p. 198-198. PMid:30042935
View Article PubMed/NCBIBaer, A.Z., et al., Prevalence of coronary artery lesions on the initial echocardiogram in Kawasaki syndrome. Archives of pediatrics & adolescent medicine, 2006. 160(7): p. 686-690. PMid:16818833
View Article PubMed/NCBIDajani, A.S., et al., Diagnosis and therapy of Kawasaki disease in children. Circulation, 1993. 87(5): p. 1776-1780. PMid:8491037
View Article PubMed/NCBIWright, V.J., et al., Diagnosis of Kawasaki Disease Using a Minimal Whole-Blood Gene Expression Signature. JAMA Pediatr, 2018. 172(10): p. e182293. PMid:30083721
View Article PubMed/NCBIKato, H., et al., Long-term consequences of Kawasaki disease. A 10- to 21-year follow-up study of 594 patients. Circulation, 1996. 94(6): p. 1379-85. PMid:8822996
View Article PubMed/NCBITaubert, K.A., A.H. Rowley, and S.T. Shulman, Nationwide survey of Kawasaki disease and acute rheumatic fever. J Pediatr, 1991. 119(2): p. 279-82. 80742-5
View ArticleMcCrindle, B.W., et al., Diagnosis, Treatment, and Long-Term Management of Kawasaki Disease: A Scientific Statement for Health Professionals From the American Heart Association. Circulation, 2017. 135(17): p. e927-e999.
Wang, Y.R. and H. Huang, Review on statistical methods for gene network reconstruction using expression data. Journal of theoretical biology, 2014. 362: p. 53-61. PMid:24726980
View Article PubMed/NCBILin, C.-C., et al., A cross-cancer differential co-expression network reveals microRNA-regulated oncogenic functional modules. Molecular biosystems, 2015. 11(12): p. 3244-3252. PMid:26448606
View Article PubMed/NCBIWang, X., et al., An R package suite for microarray meta-analysis in quality control, differentially expressed gene analysis and pathway enrichment detection. Bioinformatics (Oxford, England), 2012. 28(19): p. 2534-2536. PMid:22863766
View Article PubMed/NCBIGiulietti, M., et al., Identification of candidate miRNA biomarkers for pancreatic ductal adenocarcinoma by weighted gene co-expression network analysis. Cellular Oncology, 2017. 40(2): p. 181-192. PMid:28205147
View Article PubMed/NCBIZhang, B. and S. Horvath, A general framework for weighted gene co-expression network analysis. Statistical applications in genetics and molecular biology, 2005. 4(1). PMid:16646834
View Article PubMed/NCBIGargalovic, P.S., et al., Identification of inflammatory gene modules based on variations of human endothelial cell responses to oxidized lipids. Proceedings of the National Academy of Sciences, 2006. 103(34): p. 12741-12746. PMid:16912112
View Article PubMed/NCBICarlson, M.R., et al., Gene connectivity, function, and sequence conservation: predictions from modular yeast co-expression networks. BMC genomics, 2006. 7(1): p. 40.
Wang, X., et al., An R package suite for microarray meta-analysis in quality control, differentially expressed gene analysis and pathway enrichment detection. Bioinformatics, 2012. 28(19): p. 2534-2536. PMid:22863766
View Article PubMed/NCBISmoot, M.E., et al., Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics, 2010. 27(3): p. 431-432. PMid:21149340
View Article PubMed/NCBIKuo, H.-C., Preventing coronary artery lesions in Kawasaki disease. Biomedical journal, 2017. 40(3): p. 141-146. PMid:28651735
View Article PubMed/NCBIMakino, N., et al., Descriptive epidemiology of Kawasaki disease in Japan, 2011-2012: from the results of the 22nd nationwide survey. Journal of epidemiology, 2015: p. JE20140089. PMid:25716368
View Article PubMed/NCBIBonnert, T.P., et al., The cloning and characterization of human MyD88: a member of an IL‐1 receptor related family 1. FEBS letters, 1997. 402(1): p. 81-84. 01506-2
View ArticleArancibia, S.A., et al., Toll-like receptors are key participants in innate immune responses. Biological research, 2007. 40(2): p. 97-112. PMid:18064347
View Article PubMed/NCBIFoell, D., et al., S100A12 (EN-RAGE) in monitoring Kawasaki disease. The Lancet, 2003. 361(9365): p. 1270-1272. 12986-8
View ArticleKang, Y.J., et al., Calcineurin negatively regulates TLR-mediated activation pathways. The Journal of Immunology, 2007. 179(7): p. 4598-4607. PMid:17878357
View Article PubMed/NCBIStanislawowski, M., et al., Decreased Toll-like receptor-5 (TLR-5) expression in the mucosa of ulcerative colitis patients. J Physiol Pharmacol, 2009. 60(Suppl 4): p. 71-75.
Sharma, N., A.S. Akhade, and A. Qadri, Sphingosine‐1‐phosphate suppresses TLR‐induced CXCL8 secretion from human T cells. Journal of leukocyte biology, 2013. 93(4): p. 521-528. PMid:23345392
View Article PubMed/NCBIChamberlain, N.D., et al., TLR5, a novel and unidentified inflammatory mediator in rheumatoid arthritis that correlates with disease activity score and joint TNF-α levels. The Journal of Immunology, 2012. 189(1): p. 475-483. PMid:22661088
View Article PubMed/NCBIGohda, J., T. Matsumura, and J.-i. Inoue, Cutting edge: TNFR-associated factor (TRAF) 6 is essential for MyD88-dependent pathway but not Toll/IL-1 receptor domain-containing adaptor-inducing IFN-β (TRIF)-dependent pathway in TLR signaling. The Journal of Immunology, 2004. 173(5): p. 2913-2917. PMid:15322147
View Article PubMed/NCBIReizis, B., et al., Plasmacytoid dendritic cells: recent progress and open questions. Annual review of immunology, 2011. 29: p. 163-183. PMid:21219184
View Article PubMed/NCBIBank, S., et al., Polymorphisms in the inflammatory pathway genes TLR2, TLR4, TLR9, LY96, NFKBIA, NFKB1, TNFA, TNFRSF1A, IL6R, IL10, IL23R, PTPN22, and PPARG are associated with susceptibility of inflammatory bowel disease in a Danish cohort. PLoS One, 2014. 9(6): p. e98815. PMid:24971461
View Article PubMed/NCBILi, N., et al., LPS-induced CXCR7 expression promotes gastric Cancer proliferation and migration via the TLR4/MD-2 pathway. Diagnostic pathology, 2019. 14(1): p. 3. PMid:30636642
View Article PubMed/NCBILacronique, V., et al., A TEL-JAK2 fusion protein with constitutive kinase activity in human leukemia. Science, 1997. 278(5341): p. 1309-1312. PMid:9360930
View Article PubMed/NCBIKralovics, R., et al., A gain-of-function mutation of JAK2 in myeloproliferative disorders. New England Journal of Medicine, 2005. 352(17): p. 1779-1790. PMid:15858187
View Article PubMed/NCBICamus, G. and M. Ott, How the antiviral immune response boosts liver fat. Nature medicine, 2013. 19(6): p. 671. PMid:23744144
View Article PubMed/NCBITawfik, M.K., M.K. El-Kherbetawy, and S. Makary, Cardioprotective and anti-aggregatory effects of levosimendan on isoproterenol-induced myocardial injury in high-fat-fed rats involves modulation of PI3K/Akt/mTOR signaling pathway and inhibition of apoptosis: comparison to cilostazol. Journal of cardiovascular pharmacology and therapeutics, 2018. 23(5): p. 456-471. PMid:29685053
View Article PubMed/NCBIOshiumi, H., et al., TIR-containing adapter molecule (TICAM)-2, a bridging adapter recruiting to Toll-like receptor 4 TICAM-1 that induces interferon-β. Journal of Biological Chemistry, 2003. 278(50): p. 49751-49762. PMid:14519765
View Article PubMed/NCBIAbano, E.E., & Sam-Amoah, L. K. (2011). Effects of Different Pretreatments on Drying Characteristics of Banana Slices. Journal of Engineering and Applied Sciences, 6(3), 121-129.
Abano, E E, Owusu, J., & Engmann, F. N. (2013). Effects of ascorbic acid , salt , lemon juice , and honey on drying kinetics and sensory characteristic of dried mango. Croat. J. Food Sci. Technol, 5(2015), 1-10. PMid:26904667
View Article PubMed/NCBIAbano, Ernest Ekow. (2015). Kinetics and Quality of Microwave-Assisted Drying of Mango ( Mangifera indica ). International Journal of Food Science. PMid:26904667
View Article PubMed/NCBIAbe-Inge, V., Agbenorhevi, J. K., Kpodo, F. M., & Adzinyo, O. A. (2018). Effect of different drying techniques on quality characteristics of African palmyra palm (Borassus aethiopum) fruit flour. Food Research. .050
View ArticleAbrol, G., Devina, V., Ambika, S., & Surabhi, S. (2014). Effect of Solar Drying on Physico-chemical and Antioxidant Properties of Mango , Banana and Papaya. Natl. Acad. Sci. Lett., 37(1), 51-57. DOI:10.1007/s40009-013-0196-1
View ArticleAdepoju, L. A., & Osunde, Z. D. (2015). Quality o f Dried Banana Fruit Under Different Pretreatments a nd Drying Quality of Dried Banana Fruit Under Different Pretreatments and Drying Methods. Australian Journal of Engineering Research.
Adepoju, Lydia Abiola, & Osunde, Z. D. (2017). Effect of pretreatments and drying methods on some qualities of dried mango ( Mangifera indica ) fruit. AgricEngInt: CIGR, 19(1).
Agyapong, M. (2013). EVALUATION OF THE POST HARVEST HANDLING OF MANGO FRUIT (Mangifera indica L.) IN THE AKWAPIM-SOUTH, DANGME-WEST, LOWER MANYA AND THE YILO KROBO DISTRICTS OF GHANA. Kwame Nkrumah University of Science and Technology.
Akoy, E. O. M. (2014). Effect of Drying Temperature on Some Quality Attributes of Mango Slices. International Journal of Innovation and Scientific Research, 4(2), 91-99.
Aktas, T., Fujii, S., Kawano, Y., & Yamamoto, S. (2007). EFFECTS OF PRETREATMENTS OF SLICED VEGETABLES WITH TREHALOSE ON DRYING CHARACTERISTICS AND QUALITY OF DRIED PRODUCTS. Trans IChemE, Part C, Food and Bioproducts Processing, 85, 178-183. DOI:10.1205/fbp07037
View ArticleAl-amin, M., Hossain, S., & Iqbal, A. (2015). Effect of Pre-treatments and Drying Methods on Dehydration and Rehydration Characteristics of Carrot. Universal Journal of Food and Nutrition Science, 3(2), 23-28. DOI:10.13189/ujfns.2015.030201
Albernaz, F., Abreu, M., Roberto, J., Quintanilha, D., Hernanz, D., Heredia, F. J., … Araujo, D. L. (2017). Foam mat drying of Tommy Atkins mango : Effects of air temperature and concentrations of soy lecithin and carboxymethylcellulose on phenolic composition , mangiferin , and antioxidant capacity. Food Chemistry, 221, 258-266. DOI:10.1016/j.foodchem.2016.10.080 PMid:27979201
View Article PubMed/NCBIAOAC. (2005). Association of Official Analytical Chemists: Official methods of Analysis of AOAC International (18th ed, Vol. 2). Washington, DC.
Atungulu, G., Nishiyama, Y., & Koide, S. (2004). Respiration and climacteric patterns of apples treated with continuous and intermittent direct current electric field. Journal of Food Engineering, 63, 1-8. DOI:10.1016/S0260-8774(03)00275-9 00275-9
View ArticleBazzano, L. A., He, J., Ogden, L. G., Loria, C. M., Vupputuri, S., Myers, L., & Whelton, P. K. (2002). Fruit and vegetable intake and risk of cardiovascular disease in US adults : the first National Health and Nutrition Examination. Am J Clin Nutr, 93-99. PMid:12081821
View Article PubMed/NCBIBeveridge, T., & Weintraub, S. E. (1995). Effect of blanching pretreatment on color and texture of apple slices at various water activities. Food Research International, 28(1), 83-86. 93335-R
View ArticleBlaise, K., Ferdinand, W., Gbaha, P., & Toure, S. (2009). Mathematical modelling of the thin layer solar drying of banana , mango and cassava. Energy, 34(10), 1594-1602. DOI:10.1016/j.energy.2009.07.005
View ArticleCaparino, O. A., Tang, J., Nindo, C. I., Sablani, S. S., Powers, J. R., & Fellman, J. K. (2012). Effect of drying methods on the physical properties and microstructures of mango (Philippine 'Carabao' var.) powder. Journal of Food Engineering, 111(1), 135-148. DOI:10.1016/j.jfoodeng.2012.01.010
View ArticleDame, B., & Sahu, O. (2016). Effect of Oven Drying Temperature on Banana Powder Quality. Asian Journal of Innovative Research in Science, Engineering and Technology, 1(2), 1-8.
Dawit, A., & Dagnew, A. (2008). Banana Markets in Ethiopia. Addis Ababa, Ethiopia.
Delfiya, A., Mohapatra, D., & Kotwaliwale, N., & Mishra, A. K. (2018). Effect of microwave blanching and brine solution pretreatment on the quality of carrots dried in solar ‐ biomass hybrid dryer. Journal of Food Processing and Preservation, 42(2). DOI:10.1111/jfpp.13510
View ArticleDessalegn, Y., Assefa, H., Derso, T., & Haileslassie, A. (2016). Assessment of fruit postharvest handling practices and losses in Bahir Dar , Ethiopia. Afr. J. Agric. Res, 11(52), 5209-5214. DOI:10.5897/AJAR2016.11731
View ArticleDessalegn, Y., Assefa, H., Derso, T., & Tefera, M. (2014). Mango Production Knowledge and Technological Gaps of Smallholder Farmers in Amhara Region , Ethiopia. American Scientific Research Journal for Engineering, Technology, and Sciences, 10(1), 28-39.
Dissa, A. O., Bathiebo, D. J., Desmorieux, H., Coulibaly, O., Koulidiati, J., Claude, U., & Lyon, B. (2011). Experimental characterisation and modelling of thin layer direct solar drying of Amelie and Brooks mangoes. Energy, 36(5), 2517-2527. DOI:10.1016/j.energy.2011.01.044
View ArticleDoymaz, Ä. (2010). Effect of citric acid and blanching pre-treatments on drying and rehydration of Amasya red apples. Food and Bioproducts Processing, 88(2-3), 124-132. DOI:10.1016/j.fbp.2009.09.003
View ArticleDuresa, L. W., & Manaye, D. (2017). Phytochemical Screening and Antioxidant Activity of Selected Mango (Mangifera Indica L.) and Avocado (Persea Americana) Fruits in Illu Ababor Zone, Oromia Regional State, Ethiopia. Indo American Journal of Pharmaceutical Research., 7(09).
View ArticleFAO. (1992). Manuals of food quality control. Washington, DC, USA.
FAO. (1995). Fruit and vegetable processing. Retrieved from
View ArticleFarhana, N., Rahman, A., Shamsudin, R., Ismail, A., Nadiah, N., Karim, A., & Varith, J. (2018). Effects of drying methods on total phenolic contents and antioxidant capacity of the pomelo (Citrus grandis (L.) Osbeck) peels. Innovative Food Science and Emerging Technologies. DOI:10.1016/j.ifset.2018.01.009
View ArticleFDA. (2012). Product Specification Dried Organic Mango Slices FDFI Item code : MANO.
Fita, T. (2014). White Mango Scale, Aulacaspis tubercularis, Distribution and Severity Status in East and West Wollega Zones, Western Ethiopia. Science, Technology and Arts Research Journal, 3(3), 1-10.
View ArticleGabriel, A. A., Marie, J., Usero, C. L., Rodriguez, K. J., Diaz, A. R., & Tiangson-bayaga, C. L. P. (2015). Estimation of ascorbic acid reduction in heated simulated fruit juice systems using predictive model equations. LWT - Food Science and Technology, 64(2), 1163-1170. DOI:10.1016/j.lwt.2015.07.041
View ArticleGaloburda, R., Kruma, Z., & Ruse, K. (2012). Effect of Pretreatment Method on the Content of Phenolic Compounds , Vitamin C and Antioxidant Activity of Dried Dill. International Journal of Nutrition and Food Engineering, 6(4).
Ghiafeh, M., Vijayanand, P., Kulkarni, S. G., & Ramana, K. V. R. (2007). Effect of different pre-treatments and dehydration methods on quality characteristics and storage stability of tomato powder. LWT, 40, 1832-1840. DOI:10.1016/j.lwt.2006.12.004
View ArticleGirma, G., Garo, G., & Fetena, S. (2016). Chemical Composition of Mango ( Mangifera Indica L ) Fruit as Influence by Postharvest Treatments in Arba. Journal of Environmental Science, Toxicology and Food Technology, 10(11), 70-77. DOI:10.9790/2402-1011027077
Guiné, R. P. F. (2018). The Drying of Foods and Its Effect on the Physical-Chemical , Sensorial and Nutritional Properties. International Journal of Food Engineering, 4(2), 93-100. DOI:10.18178/ijfe.4.2.93-100
View ArticleGulzar, A., Ahmed, M., Qadir, M. A., & Imtiaz, M. (2018). Effect of Blanching Techniques and Treatments on Nutritional Quality of Dried Mango Slices During Storage. Pol. J. Food Nutr. Sci., 68(1), 5-13. DOI:10.1515/pjfns-2017-0012
View ArticleGupta, E., Purwar, S., Jaiswal, P., Chaturvedi, R., & Rai, G. K. (2016). Sensory Evaluation and Nutritional Composition of Developed Papaya-Gooseberry Jam. Food and Nutrition Sciences, 7, 600-608.
View ArticleHagos, Afework, & Zemedu, L. (2015). Determinants of Improved Rice Varieties Adoption in Fogera District of Ethiopia. Science, Technology and Arts Research Journal, 4(1), 221-228. DOI: https://doi.org/10.4314/star.v4i1.35
View ArticleHagos, Afeworki, Dibaba, R., Bekele, A., & Alemu, D. (2019). Determinants of Market Participation among Smallholder Mango Producers in Assosa Zone of Benishangul Gumuz Region in Ethiopia Determinants of Market Participation among Smallholder Mango Producers in Assosa Zone of Benishangul Gumuz Region in Ethiopia. International Journal of Fruit Science, 0(0), 1-27. DOI:10.1080/15538362.2019.1640167
View ArticleHailu, Z. and, & Worku, T. (2017). Effects of Controlled Atmosphere Storage and Temperature on Quality Attributes of Mango. Global Journal of Researches in Engineering: C Chemical Engineering Volume, 17(1).
Honja, T. (2014). Review of Mango Value Chain in Ethiopia. Journal of Biology, Agriculture and Healthcare., 4(25), 230-240.
Hota, M., Dahiya, D. S., & Kumar, S. (2017). Effect of Various Drying Methods on Drying Time and Quality of Pomegranate ( Punica granatum L .) Arils. Int.J.Curr.Microbiol.App.Sci, 6(4), 1711-1717.
View ArticleHwa, C., Lim, C., Figiel, A., Wojdyło, A., & Oziembłowski, M. (2013). Colour , phenolic content and antioxidant capacity of some fruits dehydrated by a combination of different methods. Food Chemistry, 141(4), 3889-3896. DOI:10.1016/j.foodchem.2013.06.042 PMid:23993562
View Article PubMed/NCBIIdah, P. A., & Aderibigbe, B. A. (2007). Quality Changes in Dried Tomatoes Stored in Sealed Polythene and Open Storage Systems. Leonardo Electronic Journal of Practices and Technologies, (10), 123-136.
Ikoko, J., & Kuri, V. (2007). Osmotic pre-treatment effect on fat intake reduction and eating quality of deep-fried plantain. Food Chemistry, 102, 523-531. DOI:10.1016/j.foodchem.2006.06.008
View ArticleIsmail, O. M., & Nagy, K. S. A. (2012). Characteristics of Dried Mango Slices as Affected by Pre-Treatments and Drying Type. Australian Journal of Basic and Applied Sciences, 6(5), 230-235.
Izli, N., Izli, G., & Taskin, O. (2017). Influence of different drying techniques on drying parameters of mango. Food Sci. Technol, Campinas, 37(4), 604-612.
View ArticleIzli, N., Izli, G., & Taskin, O. (2018). Impact of different drying methods on the drying kinetics , color , total phenolic content and antioxidant capacity of pineapple. CyTA - Journal of Food, 16(1), 213-221. DOI:10.1080/19476337.2017.1381174
View ArticleKabiru, A. A., Joshua, A. A., & Raji, A. O. (2013). DRYING KINETICS OF MANGO ( Mangifera Indica ). IJRRAS, 15(1), 41-50.
Kaewdam, S., Varith, J., Nitatwichit, C., & Jaturonglumlert, S. (2013). Mathematical Model of Freeze Drying on Mango. Journal of Agr. Research & Extension, 30(2), 56-67.
Karim, O. R., Awonorin, S. O., & Sanni, L. O. (2008). Effect of Pretreatments on Quality Attributes of Air-Dehydrated Pineapple Slices. Journal of Food Technology, 6(4), 158-165.
Kasso, M., & Bekele, A. (2018). Post-harvest loss and quality deterioration of horticultural crops in Dire Dawa Region , Ethiopia. Journal of the Saudi Society of Agricultural Sciences, 17(1), 88-96. DOI:10.1016/j.jssas.2016.01.005
View ArticleKidmose, U., & Martens, H. J. (1999). Changes in texture , microstructure and nutritional quality of carrot slices during blanching and freezing. Journal of the Science of Food and Agriculture, 79, 1747-1753. 1097-0010(199909)79:12<1747::AID-JSFA429>3.0.CO;2-B
View ArticleKrokida, M. K., & Maroulis, Z. B. (2001). Structural properties of dehydrated products during rehydration. International Journal of Food Science and Technology 2001, 36, 529-538.
View ArticleKumar, P. S., & Sagar, V. R. (2012). Drying kinetics and physico-chemical characteristics of Osmo- dehydrated Mango , Guava and Aonla under different drying conditions. J Food Sci Technol. DOI:10.1007/s13197-012-0658-3 PMid:25114345
View Article PubMed/NCBIKumar, S., & Shukla, R. N. (2017). Different Pre-Treatments and Storage Stability of Dehydrated Pineapple Slices. International Journal of Agricultural Science and Research, 7(2), 413-424.
Lewicki, P. P. (1998). Effect of pre ‐ drying treatment , drying and rehydration on plant tissue properties : A review. International Journal of Food Properties, 1(1), 1-22. DOI:10.1080/10942919809524561
View ArticleLink, J. V., Tribuzi, G., & Laurindo, J. B. (2017). Improving quality of dried fruits : A comparison between conductive multi-fl ash and traditional drying methods. Food Science and Technology, 84, 717-725. DOI:10.1016/j.lwt.2017.06.045
View ArticleLink, J. V., Tribuzi, G., & Laurindo, J. B. (2018). Conductive multi-flash drying of mango slices : Vacuum pulse conditions on drying rate and product properties. J Food Process Preserv. DOI:10.1111/jfpp.13440
View ArticleLiu, R. H. (2003). Health benefits of fruit and vegetables are from additive and synergistic combinations of phytochemicals. Am J Clin Nutr, 78, 3-6. PMid:12936943
View Article PubMed/NCBIMadan, A., & Pare, A. (2014). Drying and Rehydration Behaviour of Bamboo (Bambusa bambos) Shoots during Convective Tray Drying. Journal of AgriSearch, 11(3), 127-134.
Maisnam, D., Rasane, P., Dey, A., Kaur, S., & Sarma, C. (2017). Recent advances in conventional drying of foods . J Food Technol Pre, 1(1).
Maldonado, S., Arnau, E., & Bertuzzi, M. A. (2010). Effect of temperature and pretreatment on water diffusion during rehydration of dehydrated mangoes. Journal of Food Engineering, 96(3), 333-341. DOI:10.1016/j.jfoodeng.2009.08.017
View ArticleMaría, A., Inés, G., & Eduardo, C. (2012). Effect of freezing rate on quality parameters of freeze dried soursop fruit pulp. Journal of Food Engineering, 111, 360-365. DOI:10.1016/j.jfoodeng.2012.02.010
View ArticleMarques, L. G., Prado, M. M., & Freire, J. T. (2009). Rehydration characteristics of freeze-dried tropical fruits. LWT - Food Science and Technology, 42(7), 1232-1237. DOI:10.1016/j.lwt.2009.02.012
View ArticleMarques, L. G., Prado, M. M., & Freire, J. T. (2011). Vitamin C Content of Freeze-Dried Tropical Fruits. In International Congress on Engineering and Food, 22-26.
Masamba, K. G., Mkandawire, M., & Chiputula, J. (2013). Evaluation of sensory quality attributes and extent of vitamin C degradation in dried pineapple , mango and banana fruit pieces pre - treated with sodium metabisulphite and lemon juice. Glob. J. Food Sci. Technol., 1(1), 20-25.
Maskan, M. (2001). Kinetics of colour change of kiwifruits during hot air and microwave drying. Journal of Food Engineering. DOI:10.1016/S0260-8774(00)00154-0 00154-0
View ArticleMemmedov, A., Abbasov, T., & Şeker, M. (2016). Intensification of the Banana Drying Processes by Using EFL Electromagnetic Waves. World Journal of Engineering and Technology, 4, 281-289.
View ArticleMercer, D. G. (2012). A Comparison of the Kinetics of Mango Drying in Open-Air, Solar, and Forced-Air Dryers. African Journal of Food, Agriculture, Nutrition and Development, 12(7), 6835-6852.
Mezgebe, A. G., Bosha, T., & Muchie, T. D. (2016). Post-harvest losses and handling practices of durable and perishable crops produced in relation with food security of households in Ethiopia : Secondary data analysis. Journal of Stored Products and Postharvest, 7(5), 45-52. DOI:10.5897/JSPPR2016.0205
Misgana Mitiku and Tamirat Gutema. (2017). Assessment of Postharvest Handling Practices and Problems on Major Crops Assessment of Postharvest Handling Practices and Problems on Major Crops of South Ari District , South Omo Zone , SNNPR , Ethiopia. Journal of Biology, Agriculture and Healthcare, 11.
Mishra, M., Shukla, R. N., Kandasami, P., & Huirem, B. (2019). Experimental Investigation on the Effect of Pre-Treatments on Thin Layer Drying and Quality Characteristics of Green Mangoes in Forced Convection Hot Air Drying. Int.J.Curr.Microbiol.App.Sci, 8(06), 3112-3124.
View ArticleMohamed, G. F., Abdelmaguid, N. M., Ibrahim, W. A., Helmy, I. M. F., & Nadir, A. S. (2017). Effect of Different Drying Methods and Pre-Treatments on Quality Characteristics of Mango Slices. Middle East Journal of Applied Sciences, 07(03), 519-531.
Mugodo, K. (2017). Evaluation of the Effects of Pre-Drying Treatments and Drying Methods on the Drying Kinetics and Quality of Tommy Atkin Mango Slices. University of KwaZulu-Natal.
Mwamba, I., Tshimenga, K., Mulumba, L., Tshibad, C. M., & Noël, J. (2017). Comparison of two drying methods of mango ( oven and solar drying ). MOJ Food Processing & Technology Research, 5(1), 240-243. DOI:10.15406/mojfpt.2017.05.00118
View ArticleNeguse, T. B., Wanzala, F. K. R., Ali, W. M., Owino, W. O., & Mwangi, G. S. (2019). Mango ( Mangifera indica L .) production practices and constraints in major production regions of Ethiopia. African Journal of Agricultural Research Full, 14(4), 185-196. DOI:10.5897/AJAR2018.13608
View ArticleNiguse, B. (2018). Assess and Prioritize the Problems Related Postharvest Management of Horticultural Crops in Jimma Town , the Case of Bishishe Market. J Biodivers Biopros Dev, 5(1), 1-6. DOI:10.4172/2376-0214.1000168
View ArticleNtuli, V., Peter, C., Raphael, K., Tendekayi, G. H., & Jephris, G. (2017). Microbiological quality of selected dried fruits and vegetables in Maseru , Lesotho. Afr. J. Microbiol. Res, 11(5), 185-193. DOI:10.5897/AJMR2016.8130
View ArticleNyangena, I., Owino, W., Ambuko, J., & Imathiu, S. (2019). Effect of selected pretreatments prior to drying on physical quality attributes of dried mango chips. Journal of Food Science and Technology, 56(8), 3854-3863. DOI:10.1007/s13197-019-03857-9 PMid:31413411
View Article PubMed/NCBIOmolola, A. O., Jideani, A. I. O., & Kapila, P. F. (2015). Quality Properties of Fruits as Affected by Drying Operation. Critical Reviews in Food Science and Nutrition, 37-41. DOI:10.1080/10408398.2013.859563
Owolade, S. O., Akinrinola, A. O., Popoola, F. O., Aderibigbe, O. R., & Ademoyegun, O.T. and Olabode, I. A. (2017). Study on physico-chemical properties , antioxidant activity and shelf stability of carrot (Daucus carota) and pineapple (Ananas comosus) juice blend. International Food Research Journal, 24(2), 534-540.
Owureku-asare, M., & Saalia, F. K. (2014). Effect of pretreatment on physicochemical quality characteristics of a dried tomato (Lycopersicon esculentum). African Journal of Food Science, 8(5), 253-259. DOI:10.5897/AJFS2014.1156
View ArticlePerera, C. O. (2014). Drying Technology : An Selected Quality Attributes of Dried Foods. An International Journal, 23(4), 37-41. DOI:10.1081/DRT-200054180
View ArticlePott, I., Neidhart, S., Mu, W., & Carle, R. (2005). Quality improvement of non-sulphited mango slices by drying at high temperatures. Innovative Food Science and Emerging Technologies, 6, 412-419. DOI:10.1016/j.ifset.2005.05.004
View ArticlePrakash, S., Jha, S. . and, & Datta, N. (2004). Performance evaluation of blanched carrots dried by three different driers. Journal of Food Engineering, 62, 305-313. DOI:10.1016/S0260-8774(03)00244-9 00244-9
View ArticlePu, Y., & Sun, D. (2017). Combined hot-air and microwave-vacuum drying for improving drying uniformity of mango slices based on hyperspectral imaging visualisation of moisture content distribution. Biosystems Engineering, 156, 108-119. DOI:10.1016/j.biosystemseng.2017.01.006
View ArticleQuintana-Zaragoza, L., Luna-Solano, G., Salgado-Cervantes, M. A., Jim'enez-Fernandez, M., Urrea-Garcia, G. R., & Villegas-Santiago, J. (2017). OPTIMIZATION AND EXPERIMENTAL VALIDATION OF FLUIDIZED BED DRYING. Revista Mexicana de Ingenier'ıa Qu'ımica, 16(2), 625-634.
Rajkumar, P., Kulanthaisami, S., Raghavan, G. S. V., Gariépy, Y. and, & Orsat, V. (2007). Drying Kinetics of Tomato Slices in Vacuum Assisted Solar and Open Sun Drying Methods Drying Kinetics of Tomato Slices in Vacuum Assisted Solar and Open Sun Drying Methods. Drying Technology, 25(7), 1349-1357. DOI:10.1080/07373930701438931
View ArticleRwubatse, Bernard, Akubor, Peter Issah and Mugabo, E. (2014). Traditional Drying Techniques for Fruits and Vegetables Losses Alleviation in Sub-Saharan Africa. Journal of Environmental Science, Toxicology and Food Technology (IOSR-JESTFT), 8(9), 52-56. DOI:10.9790/2402-08945256
View ArticleSakagami, Y., Feroz, F., Shimizu, H., Nishioka, T., & Mori, M. (2016). Bacterial and Fungal Counts of Dried and Semi-Dried Foods Collected from Dhaka , Bangladesh , and Their Reduction Methods. Biocontrol Science, 21(4), 243-251. PMid:28003631
View Article PubMed/NCBISalazar, N. A., Alvarez, C., & Orrego, C. E. (2017). Optimization of Freezing Parameters for Freeze- Drying Mango ( Mangifera indica L .) Slices. Drying Technology An, 3937. DOI:10.1080/07373937.2017.1315431
View ArticleSalgado-, M. A., Villegas-santiago, J., Calderon-santoyo, M., & Ragazzo-sa, A. (2011). Original article Fluidized bed and tray drying of thinly sliced mango (Mangifera indica) pretreated with ascorbic and citric acid. International Journal of Food Science and Technology, 46, 1296-1302. DOI:10.1111/j.1365-2621.2011.02637.x
View ArticleScanlin, D. (1997). The Design, Construction, and Use of an Indirect, Through-Pass, Solar Food Dryer. Home Power.
Sehrawat, R., Nema, P. K., & Kaur, B. P. (2016). Effect of superheated steam drying on properties of foodstuffs and kinetic modeling. Innovative Food Science and Emerging Technologies. DOI:10.1016/j.ifset.2016.02.003
View ArticleSehrawat, R., Nema, P. K., & Kaur, B. P. (2018). Quality evaluation and drying characteristics of mango cubes dried using low-pressure superheated steam, vacuum and hot air drying methods. LWT - Food Science and Technology. DOI:10.1016/j.lwt.2018.03.012
View ArticleShah, K. A., Patel, M. B., Patel, R. J., & Parmar, P. K. (2010). Mangifera Indica ( Mango ). Pharmacognosy Reviews, 4(7). DOI:10.4103/0973-7847.65325 PMid:22228940
View Article PubMed/NCBIShi, S., Ma, X., Xu, W., Zhou, Y., Wu, H., & Wang, S. (2015). Evaluation of 28 mango genotypes for physicochemical characters , antioxidant capacity , and mineral content. Journal of Applied Botany and Food Quality, 273, 264-273. DOI:10.5073/JABFQ.2015.088.039
Shimelis Admassu, E. (2015). Postharvest sector challenges and opportunities in Ethiopia.
Singh, D., Siddiq, M., & Dolan, K. D. (2014). Total phenolics , carotenoids and antioxidant properties of Tommy Atkin mango cubes as affected by drying techniques. LWT - Food Science and Technology, 1-5. DOI:10.1016/j.lwt.2014.04.015
Soussene Boudraa, Djamel Fahloul, Driss Elothmani, Sara Zidani, M. S. (2017). Effects of Different Drying Methods on Phenols Contents and Antioxidant Activity of Azaroles (Crataegus Azarolus L.). Annals. Food Science and Technology, 18(1).
Stone, H. (2018). Example food : What are its sensory properties and why is that important ? Npj Science of Food, 2(11). DOI:10.1038/s41538-018-0019-3 PMid:31304261
View Article PubMed/NCBISurendar, J., Dm, S., & Pd, S. (2018). Effect of drying on quality characteristics of dried tomato powder. Journal of Pharmacognosy and Phytochemistry, 7(2), 2690-2694.
Tadesse, F. T., Abera, S., & Solomon, W. K. (2016). Rehydration Capacity and Kinetics of Solar-Dried Carrot ( Daucus carota ) Slices as Affected by Blanching and Osmotic Pretreatments. Int. J. Food Eng., 12(2), 203-210. DOI:10.1515/ijfe-2015-0210
View ArticleTewari, S., Sehrawat, R., Nema, P. K., & Pal, B. (2016). Preservation e ff ect of high pressure processing on ascorbic acid of fruits and vegetables : A review. Journal of Food Biochemistry 2016;, (August 2015), 1-14. DOI:10.1111/jfbc.12319
View ArticleVelickova, E., Winkelhausen, E., & Kuzmanova, S. (2013). Physical and sensory properties of ready to eat apple chips produced by osmo-convective drying. J Food Sci Technol. DOI:10.1007/s13197-013-0950-x PMid:25477635
View Article PubMed/NCBIWakjira, M. (2010). Solar drying of fruits and windows of opportunities in Ethiopia. African Journal of Food Science Vol 4.(13) Pp. 790 - 802 Available Online Http://Www.Academicjournals.Org/Ajfs ISSN, 4(December), 790-802.
Wang, H., Fu, Q., Chen, S., Hu, Z., & Xie, H. (2018). Effect of Hot-Water Blanching Pretreatment on Drying Characteristics and Product Qualities for the Novel Integrated Freeze-Drying of Apple Slices. Journal of Food Quality.
View ArticleWang, Q., Li, S., Han, X., Ni, Y., Zhao, D., & Hao, J. (2019). Quality evaluation and drying kinetics of shitake mushrooms dried by hot air, infrared and intermittent microwave-assisted drying methods. LWT - Food Science and Technology. DOI:10.1016/j.lwt.2019.03.020
View ArticleWHO. (2003). Fruit and vegetable promotion initiative. Geneva, Switzerland.
WHO. (2019). Fruits and vegetables are important components of a healthy diet. Retrieved from
View ArticleWiersinga, R. C. (2009). Business opportunities in the Ethiopian fruit and vegetable sector. LEI Wageningen UR, The Hague.
Wilson, R. A., Kadam, D. M., Chadha, S., & Sharma, M. (2012). Foam Mat Drying Characteristics of Mango Pulp. International Journal of Food Science and Nutrition Engineering, 2(4), 63-69. DOI:10.5923/j.food.20120204.03
View ArticleWitthuhn, R. C., Engelbrecht, S., Joubert, E., & Britz, T. J. (2005). Microbial content of commercial South African high-moisture dried fruits. Journal of Applied Microbiology, 98, 722-726. DOI:10.1111/j.1365-2672.2004.02500.x PMid:15715876
View Article PubMed/NCBIXiao, H.-W., Yao, X.-D., Lin, H., Yang, W.-X., & Gao, J.-S. M. and Z.-J. (2011). Effect of SSB (Superheated Steam Blanching) Time and Drying Temperature on Hot Air Impingement Drying Kinetics and Quality Attributes of Yam Slices. Journal of Food Process Engineering, 1-21. DOI:10.1111/j.1745-4530.2010.00594.x
View ArticleYan, W., Zhang, M., Huang, L., Tang, J., Mujumdar, A. S., & Sun, J. (2010). Studies on different combined microwave drying of carrot pieces. International Journal of Food Science and Technology, 45, 2141-2148. DOI:10.1111/j.1365-2621.2010.02380.x
View ArticleYi, J.-Y., Lyu, J., Jin-Feng, Zhou, L.-Y., & Zhou, M. (2017). Hot air drying and freeze drying pre-treatments coupled to explosion pu ffi ng drying in terms of quality attributes of mango , pitaya , and papaya fruit chips. J Food Process Preserv., 1-10. DOI:10.1111/jfpp.13300
View ArticleYoon, J., & Kim, B. (2012). Lab-on-a-Chip Pathogen Sensors for Food Safety. Sensors, 12, 10713-10741. DOI:10.3390/s120810713 PMid:23112625
View Article PubMed/NCBIYusufe, M., Mohammed, A. and, & Satheesh, N. (2017). Effect of Drying Temperature and Duration on Nutritional Quality of Cochoro Variety Tomato (Lycopersicon Esculentum L.). Annals. Food Science and Technology, 18(2).
Zhao, J., Liu, F., Wen, X., Xiao, H., & Ni, Y. (2015). State diagram for freeze-dried mango : Freezing curve , glass transition line and maximal-freeze-concentration condition. JOURNAL OF FOOD ENGINEERING, 157(2015), 49-56. DOI:10.1016/j.jfoodeng.2015.02.016
View ArticleZou, K., Teng, J., Huang, L., Dai, X., & Wei, B. (2013). Effect of osmotic pretreatment on quality of mango chips by explosion puf fi ng drying. LWT - Food Science and Technology, 51(1), 253-259. DOI:10.1016/j.lwt.2012.11.005
View Article