Ajey Kumar Pathak
Scientist, ICAR-National Bureau of Fish Genetic Resources
Email: pathakajey@gmail.com, Ajey.Pathak@icar.gov.in
© 2019 Sift Desk Journals. All Rights Reserved
VOLUME: 3 ISSUE: 2
Page No: 261-268
Ajey Kumar Pathak
Scientist, ICAR-National Bureau of Fish Genetic Resources
Email: pathakajey@gmail.com, Ajey.Pathak@icar.gov.in
Mahender Singh Verma, Ajey Kumar Pathak*, Anshul Tiwari, Murali S., Ravindra Kumar, Naresh Sahebrao Nagpure, Iliyas Rashid, Rameshwar Pati and Basdeo Kushwaha
ICAR-National Bureau of Fish Genetic Resources, Canal Ring Road, P.O. Dilkusha, Lucknow - 226 002 INDIA
Maria Hrmova(maria.hrmova@adelaide.edu.au)
Ajey Kumar Pathak, Interaction profiling of growth hormone receptor and growth hormone proteins in Labeo rohita: an insight using molecular modelling and docking approaches(2019)Journal of Computational Chemistry & Molecular Modeling 3(2)p:261-268
To elicit growth and related functions, the interaction of growth hormone receptor (GHR) with growth hormone (GH) is vital as this contact passes the message downstream to the metabolic pathways. The crystal structure of GHR and GH in Labeo rohita is still vague. This study discusses prediction of 3D structures of GHR and GH, validation of predicted structures and binding affinity of GHR with GH. Discovery studio 4.1 software was used for homology modelling and docking of GHR with GH. The best model for GHR (DOPE score: -16527.1) and GH (DOPE score: -17613.3) were used for docking by ZDOCK pro. The docked complex obtained from the interaction of GHR with GH (GHR-GH, ZDOCK score: 19.26) was refined by RDOCK (RDOCK score: -25.642). The findings revealed that GHR-GH complex is stable and interacting amino acids in GHR are R43, E44, I108, D126, E127, D164, R159, W166, W169, and in GH are F3, L45, K49, P61, R64, D116, G128, K172, T175, R178. The interaction of amino acids in the active pockets of complex and its duration provides insight into the actions of GH in Labeo rohita. In addition, docking of GHR with GH provides information about the type of interaction at the binding sites of ligand and receptor and interacting amino acids viz. PHE3, ALA22, TYR98, LEU114, MET145, VAL149, LEU150 and PHE199. Thus, docking of the GHR with GH is useful in understanding the structural information and developing specific receptor antagonists beside providing support in establishing therapeutic approaches in teleost fishes.
Keywords: Growth hormone; Growth hormone receptor; Homology modelling; Labeo rohita; Protein–protein interaction; Structure prediction
Growth hormone (GH) is a pituitary protein that has pleiotropic effects on fish, including growth, development, immune function, osmoregulation, sexual maturation, reproduction, appetite regulation, starvation, protein and lipid metabolism [1-4]. Growth hormone activates the Janus kinase (JAK)–signal transducer and activator of transcription (STAT) signalling pathway. The recent studies have provided a novel understanding on the mechanism of JAK2 activation by growth hormone binding to its receptor. GH protein of Labeo rohita is a 4-helix bundle protein comprised of 207 amino acid residues. This protein has 23.521 kDa molecular weight, 6.74 isoelectric point and positions of glycosylation sites are 130 and 182. GH protein holds four cysteine residues that generate the functional 3D conformation of the hormone by creating two intramolecular disulfide (S-S) bonds in all the vertebrates. In Labeo rohita, cysteine residues are present in the GH protein at positions 49, 158, 175 and 183 respectively [5]. Again, it has an additional conjoint Cysteine at position 120, whose usefulness has never been identified [6]. Additionally, adjustment of cysteine to Alanine influences refolding of GH, lesser affinity in cellular response, receptor binding and natural processes [7].
The Growth Hormone Receptor (GHR) belongs to tyrosine-kinase linked receptor of Class-I cytokine receptor super family [8,9]. GHR is 604 amino acid long protein of 67.207 kDa molecular weight and contains an extracellular N-terminal domain, a transmembrane domain and an intracellular domain [10]. When GH interacts with GHR, the interaction passes message downstream to the metabolic pathways for stimulating growth and other functions. One GH molecule binds with one GHR molecule on the cell surface of the target tissue and the second molecule of GHR dimerises with GHR-GH complex and therafter intracellular signalling process starts [11]. This interaction of one GH with two receptor molecules triggers the GHR associated Janus kinase 2 (JAK2). This stimulation initiates the phosphorylation of the receptor tyrosine and permits for the recruitment of a class of highly related cytoplasmic proteins termed as signal transducers and activators of transcription (STATs) [12]. The GHR bound STATs are phosphorylated by JAK2 [13]. After phosphorylation, the cytoplasmic STATs form dimmers, which translocate in the nucleus and bind to a palindromic DNA motif present in the STAT inducible gene promoters. This binding initiates the transcription of target genes [14].
Labeo rohita is an important aquaculture species in the Indian subcontinent and the prolonged interaction of GHR and GH can be important in increasing the growth of this species. Till dated, the experimental 3D structures of GHR and GH in L. rohita are not present in the Protein Databank (PDB). Therefore, it is imperative to predict the tertiary structure of targeted proteins to understand its mode of action. In many regulatory processes, the biomolecular interactions have a central role in controlling cellular actions and often considered as a concern of specific protein–protein contacts [15]. Pathway regulations, signal transduction, cell cycle proliferation, cellular architecture biosynthetic and immune response etc. are few vital processes administered by the protein-protein interactions. The cellular and molecular level understandings of these actions can be well known by the quantitative structural and functional analysis of the interacting protein complexes [16,17]. The present insilico study will help in the genetic manipulation of GH/ GHR for improved growth in Labeo rohita.
Homology search
The protein sequences of GHR (acc no. UniProtKB/Swiss-Prot: Q5Y4D3) and GH (acc no. UniProtKB/Swiss-Prot: Q9W6J7) of Labeo rohita were retrieved from UniProt-KB [18]. The sequences of the GHR and GH were downloaded to find the related protein structures using BLAST [19] in the RCSB-PDB [20]. The associated protein structures of high alignment score were selected as template.
Model building of GHR and GH
On the premise of alignment of target with template sequences of the GHR and GH, 3D homology models of the targeted proteins were predicted from the crystal structural coordinates of the templates through Discovery Studio 4.1.
Validation of GHR and GH model
The 3D structural models of GHR and GH were validated using PROCHECK [21], Verify 3D [22] and PROSA [23]. The vailadation of these models was done to ensure the correctness of the fold/structure, find errors over confined regions and look into the stereochemical properties such as bond angles and lengths.
Docking of GHR with GH and refinement of the docked complex
To address the protein-protein docking, a number of docking algorithms have been proposed such as FTDock, DOT, and GRAMM [24-27]. For docking of GHR with GH, ZDOCK of Discover Studio 4.1 software using ZRANK (ranks docked poses) was used. ZDOCK is a rigid body docking program and uses a simple shape complementarity method called Pairwise Shape Complementarity (PSC) [27]. ZDOCK is an inflexible body docking procedure that includes Fast Fourier Transform (FFT) and an extensive six-dimensional search in the translational and rotational space between the two molecules [28]. The RDOCK procedure in Discovery Studio 4.1 was used to refine the docked poses [27,29]. The interaction between GHR and GH was determined in two steps that involved preliminary protein-protein docking followed by refinement of the docked complex. In the first step, putative binding poses were predicted using ZDOCK with 6 rotational sampling and randomization that provided 2000 best docked poses. These conformations were ranked using ZRank, a scoring function that customs an amalgamation of electrostatics and desolvation parameters with PSC [30]. In the next step, 18 topmost scoring poses out of 2000 poses were used for energy minimization using the CHARMM force field [31]. The RDOCK procedure refined the docked complex by optimizing the polar and charge interaction. It provided the electrostatic (Eelec) and desolvation (∆GACE) energies of the refined structure by removing clashes. The desolvation energy of the protein complex is the sum of the atomic contact energy (ACE) of total receptor-ligand atom couples inside the threshold of 6 Å. The binding affinity (∆Gbinding) calculated using RDOCK scoring function between the protein complexes is the sum of desolvation and electrostatic involvements.
∆Gbinding = ∆GACE + β X ∆Gelec
where β is a scaling factor and a value of 0.9 was used in the present study.
Homology modelling
The homology searches provided 3HHR(B) template for GHR and 1HWG(A) for GH. The identity between GHR and 3HHR(B) sequences was found 48%. Similarly, the identity between GH and 1HWG(A) sequences was found 36%. A structure based alignment between GHR and GH was done with these templates.
3D model building for GHR and GH and refinement
The 3D structure of GHR and GH in Labeo rohita were predicted using homology search through 3D structure of human GHR and GH (3HHR and 1HWG). The score obtained from each sequence alignment was used to build models using the homology modelling of Discovery Studio 4.1. The MODELLER auto model [32] was used to predict 3D structures of target proteins and 100 structures were predicted for both. Out of 100 structures, the best predicted structures for GHR and GH were selected based on the lowest value in the Probability Density Functions (PDF), Total Energy, PDF Physical Energy and Discrete Optimized Protein Energy (DOPE). The predicted structures of the GHR and GH were Q5Y4D3.M0069 (PDF Total Energy= 2374.88; PDF Physical Energy = 685.856; DOPE Score = - 16527.1) and Q9W6J7.M0019 (PDF Total Energy= 716.264; PDF Physical Energy = 396.458; DOPE Score = - 17613.3) respectively. These are best models of GHR (Fig. 1a) and GH (Fig. 2a) compared to other predicted models. The extracellular domain of the GHR contains 2 fibronectin type III (FNIII) subdomains. The first FNIII domain spans amino acids 41–135 and the second domain spans 137–240 amino acids. The GH model (Q9W6J7.M0019) is 4 a-helix bundle made of hydrophobic amino acids, arranged in an unusual up–up–down–down topology distinguishing the hematopoietic factors’ family of peptides. The 3D structure of GH is stabilized by 2 disulfide bonds in addition to numerous hydrogen bonds among different amino acid residues. The structural properties of GHR and GH are almost similar to the human’s growth hormone receptor and growth hormone [33,34].
Analysis of predicted models of GHR and GH
The stereochemical analysis of the predicted models of the GHR and GH using PROCHECK provided the Ramachandran plot presented in Fig. 1b and 2b respectively. The plot shows 180 amino acid residues in GHR and 188 in GH, which indicates good distributions of residues. In GHR, out of 180, 88.3% amino acid residues are in the most favoured and allowed regions. Only one amino acid residue is present in the generously allowed region. In GH, out of 188, 93.8% amino acid residues are in the most favoured and allowed regions and three amino acid residues are in the generously allowed region. The associated problem with the predicted 3D structure of GHR and GH was to validate and determine the accurate model. The modification of numerous protein structures for précising errors has stimulated for expansion of the new principles. Therefore, it is imperative to review the rationality of X-ray and Nuclear Magnetic Resonance (NMR) structures in addition to creating energetic and empirical procedures for assessing the accuracy of protein structures. As Verify3D evaluates the compatibility of an atomic model (3D) with its own amino acid sequence (1D), the analysis of GHR and GH using Verify 3D provided consistency with their respective amino acids having 95.5% and 27.6% residues. The assessment of GHR and GH models was done on the basis of the quality of the predicted structures (Q5Y4D3.M0069 and Q9W6J7.M0019). The path with the best Root Mean Square Deviation (RMSD) of GHR and GH models was subjected to the dynamic programming to attain an optimal alignment. RMSD of these models having values 3.61 and 1.02 provided 88.3% and 93.8% accuracies respectively. The PROSA analysis of GHR and GH, which is a measure of statistical impact of the result relative to alignment of the random structures revealed values of z-score -4.01 and -3.69 respectively (Fig. 1c and 2c). Review of literatures has evidenced that proteins with a similar fold should have z-score -3.5 or better. Thus, predicted models of GHR and GH are worth for interaction studies.
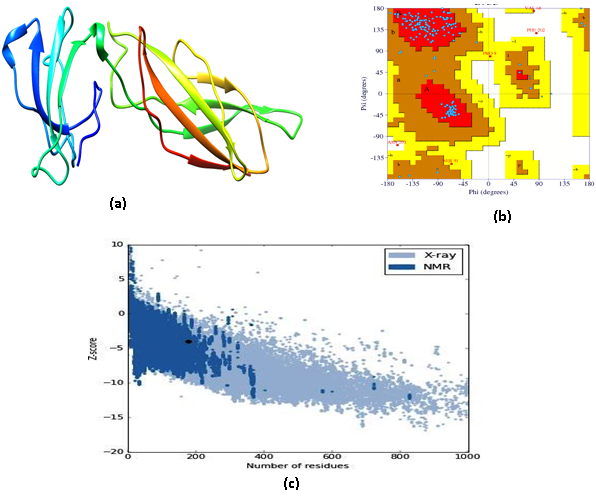
Fig. 1 (a)The best predicted model of GHR protein (Q5Y4D3.M0069); (b) Ramachandran Plot of the GHR protein model calculated by PROCHECK; (c) ProSA analysis of modeled GHR protein structure
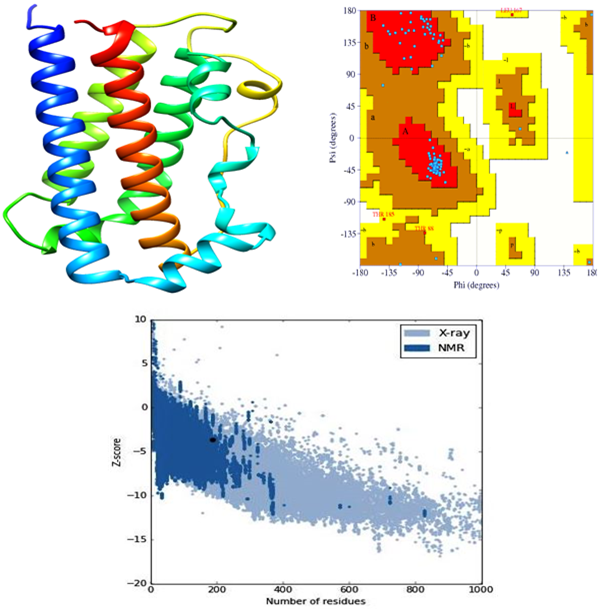
Fig. 2 (a) The best predicted model of GH protien (Q9W6J7.M0019); (b) Ramachandran Plot of the GH protein model calculated by PROCHECK; (c) ProSA analysis of modeled GH protein structure
Docking analysis of GHR with GH and refinement of docked complex
The protein-protein docking method was applied for obtain the potential structures of GHR-GH complex and only a few of them resembled the native structure. 2000 protein poses per job were predicted by ZDOCK and these poses were subjected for filtration. Afterward, the flexible energy minimization procedure (RDOCK) was applied for ranking and refinement of the predicted poses and 18 best poses were selected. The interaction analysis using ZDock and RDock revealed that GHR binds with GH with high affinity. The best pose (Pose585) of the predicted GHR-GH complex conformations was selected for analyzing the binding and interacting amino acid residues. The analysis illustrating the accurate binding mode of GHR and GH are depicted in Table 1 and Fig. 3. Fig. 3 clearly shows the consistent intermolecular hydrogen bonds created on the active sites of GHR and GH complex. Thus, GHR-GH complex is stable. The interacting amino acid residues in GHR are R43, E44, I108, D126, E127, D164, R159, W166, W169 and in GH are F3, L45, K49, P61, R64, D116, G128, K172, T175, R178. Besides, docking of GHR with GH provides information about the type of interaction at the binding sites of ligand and receptor and interacting amino acids viz. PHE3, ALA22, TYR98, LEU114, MET145, VAL149, LEU150 and PHE199. These amino acid residues also reported in GHR-GH complex of human [35-36] contribute the overall stability of the complex.
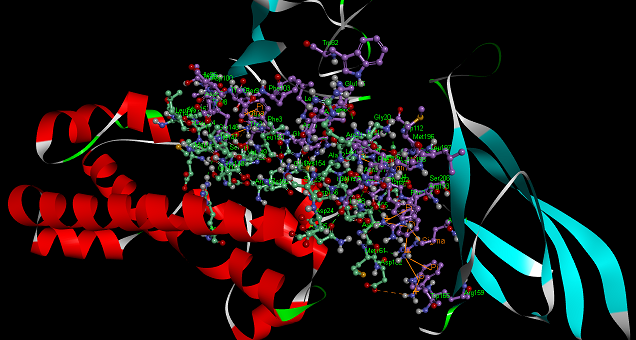
Fig. 3 Top ranked GHR-GH complex pose using ZDOCK (Purple color shows receptor interacting residues and green color shows ligand interacting residues)
Table 1 Growth hormone (GH) interaction energy calculation with Growth hormone Receptor (GHR) of Labeo rohita
|
Receptor |
Ligand |
Pose Number |
ZDock Score |
ZRank Score |
E_vdw1 |
E_Vdw 2 |
E_ele c2 |
E-sol |
E_RDock |
|
Growth hormone receptor (Ghr) |
Growth hormone (Gh) |
Pose585 |
19.26 |
5.584 |
-93.405 |
-116.342 |
-29.393 |
0.8 |
-25.642 |
The 3D structures of the GHR and GH provides the mechanism to understand about GH in action and its native conformations. Moreover, the 3D structure prediction by comparative modeling provides an alternative approach to visualise the structural facts about the new proteins by comparing with the proteins, the structure of which has been determined by X-ray crystallography or NMR. The protein-protein docking revealed the binding affinity of GHR with GH and its orientation. This study provides the information on amino acid residues contributing in interaction at the binding sites of ligand and receptor. The extracellular domain of the GHR that directly interacts with GH contains 2 fibronectin type III (FNIII) subdomains. GHR-GH complex also showed the interacting amino acid residues in GHR namely R43, E44, I108, D126, E127, R159, D164, W166 and W169 lies in these two FNIII subdomains. During interaction with GHR, the whole GH interacts with extracellular domain of GHR and thereby reveals the interacting amino acids namely F3, L45, K49, P61, R64, D116, G128 K172, T175, R178 are employed from whole GH polypeptide of 207 amino acids. Besides, docking of GHR with GH provides information about the type of interaction at the binding sites of ligand and receptor and interacting amino acids viz. PHE3, ALA22, TYR98, LEU114, MET145, VAL149, LEU150 and PHE199. These amino acid residues are also reported in Human GHR-GH complex and contribute the overall stability of the complex. These interacting amino acids are the potential sites for targeted genome engineering using tools like CRISPR/Cas (clustered regularly interspaced short palindromic repeats/CRISPR-associated protein). The ultimate impact of GH i.e. the growth of individual organism depends on the binding duration of GHR and GH. Thus, the stronger and longer binding may provide stronger and prolonged signals down the cascade and elicit better growth. With the advancement of genome engineering tools, there is possibility of substitution of a few of these interacting amino acids that may provide better complex formation and growth. Further studies have shown that GH binding to GHR results in activation of JAK2, which subsequently phosphorylates multiple tyrosine residues on the inter cellular domain (ICD) of the receptor [37]. Upon GH binding, the structural change occurs in the extracellular domain that changes the receptor transmembrane domain -from a parallel form to a left-handed crossover form. Thus, the GH has wide range of actions in regulation of metabolism especially in liver and adipose tissue besides actions on immune and reproduction [38]. Moreover, when GHR interact with Janus kinase, signal transducers and transcription activators it stimulates metabolic effects and insulin-like growth factor (IGF) synthesis [39]. Thus, docking of the GHR and GH provides insights in understanding the structural information and developing specific receptor antagonists. Additionally, this may support in establishing the therapeutic approaches in teleost fishes.
Authors are thankful to the Director of ICAR-National Bureau of Fish Genetic Resources, Lucknow, Uttar Pradesh, India for providing the necessary facilities to carry out the work supported by Centre for Agricultural Bioinformatics (CABin) Scheme of ICAR-Indian Agricultural Statistics Research Institute, New Delhi, India established under Indian Council of Agricultural Research (ICAR), Ministry of Agriculture and Farmer Welfare, Govt. of India, New Delhi.
Author's contribution
MSVand NSN conceived the concept and edited the manuscript. AKP and AKT did all the molecular works and drafted the manuscript. AKP edited the manuscript also. MS, IR, RP and BK supported the work.
Lia WS, Chen D, Wong AOL, Lin HR (2005) Molecular cloning, tissue distribution, and ontogeny of mRNA expression of growth hormone in orange-spotted grouper (Epinephelus coioides). General and Comparative Endocrinology 144(1):78-89. PMid:16055124
View Article PubMed/NCBIPoppinga J, Kittilson J, Cormick SD, Sheridan MA (2007) Effects of somatostatin on the growth hormone-insulin-like growth factor axis and seawater adaptation of rainbow trout (Oncorhynchus mykiss). Aquaculture 273: 312-319.
View ArticleLeedom TA, Uchida K, Yada T, Richman NH, Byatt JC, Collier RJ, Hirano T, Grau EG (2002) Recombinant bovine growth hormone treatment of tilapia: growth response, metabolic clearance, receptor binding and immunoglobulin production. Aquaculture 207:359-380. 00767-0
View ArticleYun L, Xiaochun L, Yong Z, Pei Z, Haoran L (2007) Molecular cloning, characterization and distribution of two types of growth hormone receptor in orange-spotted grouper (Epinephelus coioides). General and Comparative Endocrinology 152:111-122. PMid:17382945
View Article PubMed/NCBIVenugopal T, Mathavan S, Pandian TJ (2002) Molecular cloning of growth hormone encoding cDNA of Indian major carps by a modified rapid amplification of cDNA ends strategy. Journal of Biosciences 27(3): 261-272.
View ArticleWaters M J, Brooks A J, Chhabra Y (2014) A new mechanism for growth hormone receptor activation of JAK2, and implications for related cytokine receptors. JAK-STAT. 3, e29569. PMid:25101218
View Article PubMed/NCBIRajesh R, Majumdar KC (2008) The growth hormone-encoding gene isolated and characterized fromLabeo rohitaHamilton is expressed in CHO cells under the control of constitutive promoters in 'autotransgene' constructs. Fish Physiol Biochem 34:413-436. PMid:18958599
View Article PubMed/NCBICalduch-Giner J, Duval H, Chesnel F, Boeuf G, Pérez-Sánchez J, Boujard D (2001) Fish growth hormone receptor: molecular characterization of two membrane-anchored forms. Endocrinology 142(7):3269-3273. PMid:11416050
View Article PubMed/NCBIBenedet S, Johansson V, Sweeney G, Burgos MG, Bjornsson BT (2005) Cloning of two Atlantic salmon growth hormone receptor isoforms and in vitro ligand-binding response. Fish Physiology and Biochemistry 31:315-329.
View ArticleJiao B, Huang X, Chan Chi B, Zhang L, Wang D, Cheng CHK (2006) The co-existence of two growth hormone receptors in teleost fish and their differential signal transduction, tissuedistribution and hormonal regulation of expression in seabream. Journal of Molecular Endocrinology 36:23-40. PMid:16461924
View Article PubMed/NCBIDeng L, Zhang WM, Lin HR, Christopher HKC (2004) Effects of food deprivation on expression of growth hormone receptor and proximate composition in liver of black seabream Acanthopagrus schlegeli. Comparative Biochemistry and Physiology Part B. 137:421-432. PMid:15081994
View Article PubMed/NCBIBrooks A J, Waters M J (2010) The growth hormone receptor: mechanism of activation and clinical implications. Nature Reviews Endocrinology. 6:515-525. PMid:20664532
View Article PubMed/NCBIHerrington J, Carter-Su C (2001) Signalling pathways activated by the growth hormone receptor. Trends Endocrinol Metab. 12:252-257. 00423-4
View ArticleMarins LF, Iyengar A, Maclean N, Levy JA, Sohm F (2002) Simultaneous overexpression of GH and STAT5b genes inhibits the STAT5 signalling pathway in tilapia (Oreochromis niloticus) embryos. Genetics and Molecular Biology 25(3):293-298.
View ArticleEisenberg D, Marcotte E M, Xenarios I, Yeates T O (2000) Protein function in the post-genomic era. Nature405:823-826. PMid:10866208
View Article PubMed/NCBIBrooks AJ, Rebecca AP, Olivier G, Kathryn AT, Kristopher MB, Craig JM, et al. (2014) Mechanism of Activation of Protein Kinase JAK2 by the Growth Hormone Receptor. Science 344:1249783. PMid:24833397
View Article PubMed/NCBIWaters MJ (2016) The growth hormone receptor. Growth Hormone & IGF Research 28:6-10. PMid:26059750
View Article PubMed/NCBIBoutet E, Lieberherr D, Tognolli M, et al., (2016) UniProtKB/Swiss-Prot, the Manually Annotated Section of the UniProt KnowledgeBase: How to Use the Entry View. Methods Molecular Biology 1374:23-54. PMid:26519399
View Article PubMed/NCBIAltschul SF, Gish W, Miller W, Myers EW, Lipman, DJ (1990) Basic local alignment search tool. Journal of Molecular Biology 215:403-410. 80360-2
View ArticleBerman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The Protein Data Bank. Nucleic Acids Research 28:235-242. PMid:10592235
View Article PubMed/NCBILaskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK - a program to check the stereochemical quality of protein structures. Journal of Applied Crystallography 26:283-291.
View ArticleEisenberg D, Lüthy R, Bowie JU (1997) VERIFY3D: assessment of protein models with three-dimensional profiles. Methods in enzymology 277:396-404. 77022-8
View ArticleWiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Research. 35:407-410. PMid:17517781
View Article PubMed/NCBIGabb HA, Jackson RM, Sternberg MJE (1997). Modelling protein docking using shape complementarity, electrostatics and biochemical information. Journal of Molecular Biology 272:106-120. PMid:9299341
View Article PubMed/NCBIMandell JG, Roberts VA, Pique ME, Kotlovyi V, Mitchell JC, Nelson E, Tsigelny I, Ten Eyck LF (2001). Protein docking using continuum electrostatics and geometric fit. Protein Eng.14:105-113.
View ArticleVakser IA (1997). Evaluation of GRAMM low-resolution docking methodology on the hemagglutinin-antibody complex, Proteins, Suppl.1:226-230. (GRAMM performance at CASP). 1097-0134(1997)1+<226::AID-PROT31>3.3.CO;2-0
View ArticleChen R, Li L, Weng Z (2003) ZDOCK: an initial-stage protein-docking algorithm. Proteins 52(1):80-87. PMid:12784371
View Article PubMed/NCBIHarrison R W, Kourinov I V, Andrews L C (1994) The Fourier-Greens function and the rapid evaluation of molecular potentials. Protein Eng. 7:359-369. PMid:8177885
View Article PubMed/NCBIRuiz-Carmona S, Alvarez-Garcia D, Foloppe N, Garmendia-Doval AB, Juhos S, et al. (2014) rDock: A Fast, Versatile and Open Source Program for Docking Ligands to Proteins and Nucleic Acids. PLoS Comput Biol 10(4): e1003571. PMid:24722481
View Article PubMed/NCBIPierce B, Weng Z (2007) ZRANK: reranking protein docking predictions with an optimized energy function. Proteins 67(4):1078-86. PMid:17373710
View Article PubMed/NCBIVanommeslaeghe K, Hatcher E, Acharya C, et al. (2010) CHARMM General Force Field (CGenFF): A force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. Journal of computational chemistry31(4):671-690. PMid:19575467
View Article PubMed/NCBIWebb B, Sali A (2014) Comparative Protein Structure Modeling Using Modeller. Current Protocols in Bioinformatics, John Wiley & Sons, Inc., 5.6.1-5.6.32. PMid:25199792
View Article PubMed/NCBIDe Vos AM, Ultsch M, Kossiakoff AA (1992) Human growth hormone and extracellular domain of its receptor: Crystal structure of the complex. Science255:306-312. PMid:1549776
View Article PubMed/NCBIFuh G, Mulkerrin MG, Bass S, McFarland N, Brochier M, Bourell JH, et al. (1990) The human growth hormone receptor. Secretion from Escherichia coli and disulfide bonding pattern of the extracellular binding domain. J Biol Chem. 265:3111-3115.
Cunningham BC, Wells JA (1993) Comparison of a structural and a functional epitope. Journal of Molecular Biology234:554-563. PMid:7504735
View Article PubMed/NCBIBass SH, Mulkerrin MG, Wells JA (1991) A systematic mutational analysis of hormone-binding determinants in the human growth hormone receptor. Proc Natl Acad Sci U S A. 88:4498-4502. PMid:2034689
View Article PubMed/NCBIDehkhoda F, Lee CMM, Medina J, & Brooks A J (2018) The Growth Hormone Receptor: Mechanism of Receptor Activation, Cell Signaling, and Physiological Aspects. Frontiers in Endocrinology, 9, 35. . https://doi.org/10.3389/fendo.2018.00035 PMid:29487568
View Article PubMed/NCBIWaters M J (2016) The growth hormone receptor. Growth Hormone & IGF Research, 28, 6-10. PMid:26059750
View Article PubMed/NCBILin S, Li C, Li, C, & Zhang X (2018) Growth Hormone Receptor Mutations Related to Individual Dwarfism. International journal of molecular sciences, 19(5), 1433. PMid:29748515
View Article PubMed/NCBI