Sadik BUYUKYORUK,
Tel.: +902562470700
E-mail: sbuyukyoruk@adu.edu.tr
© 2019 Sift Desk Journals. All Rights Reserved
VOLUME: 4 ISSUE: 5
Page No: 770-776
Sadik BUYUKYORUK,
Tel.: +902562470700
E-mail: sbuyukyoruk@adu.edu.tr
Sadik BUYUKYORUK1, Devrim BEYAZ1, Ergun Omer GOKSOY1, Filiz KOK1, Pelin KOCAK1, Nevin TUZCU2, Ozlem Sahan YAPICIER3
1Adnan Menderes University, Faculty of Veterinary Medicine, Department of Food Hygiene and Technology, Isikli/Aydın/Turkey;
2Cumhuriyet University, Faculty of Pharmacy, Department of Basic Pharmaceutical Sciences Sivas/ Turkey;
3Mehmet Akif Ersoy University, Faculty of Veterinary Medicine, Department of Microbiology, Burdur/ Turkey
Palmiro Poltronieri(palmiro.poltronieri@ispa.cnr.it)
Anna Kristina Witte(Anna.witte@vetmeduni.ac.at)
Tong Zhao(tongzhao@uga.edu)
Spiros Paramithiotis(sdp@aua.gr)
Sadik Buyukyoruk, Survival of Listeriamonocytogenes during the production and ripening of Boza(2019)SDRP Journal of Food Science & Technology 4(5)
This study intended to determine the survival of Listeria monocytogenes during the production and fermentation of the traditional Turkish wheat-based beverage, Boza. L. monocytogenes was inoculated at a level of 5.70 log cfu/ml and survival at 30°C was studied during Boza fermentation. The study also determined the pH changes during that period. The results indicated that L. monocytogenes count tended to decrease rapidly together with a sharp decrease in the pH of boza after 48th hour storage. Additionally, L. monocytogenes was not detected after 72 hour of boza production. It was concluded that this result was attributed by the low pH formation during boza fermentation as well as the antilisterial activity of the Enterococcus faecalis and Enterococcus faecium strains that were isolated from the boza as the dominant species.
Keywords: Boza; Enterococcus spp; Listeria monocytogenes; ribotyping; survival.
Boza is a beverage prepared by adding drinking water to one or more types of purified grits or flour from different cereal varieties (e.g. proso millet, rice, wheat, corn). In the traditional method, this procedure is followed by cooking and adding sugar. Finally, alcohol and lactic acid fermentations are applied to end the preparation. It has been reported that proso millet is the most favorable ingredient for the ideal boza (Anonymous, TS 9778, 1992; Arici and Daglioglu, 2002).This fermentative beverage is mostly consumed in Balkan countries; being produced by the fermentation of lactic acid bacteria (LAB) (Lactobacillus, Lactococcus, Pediococcus and Leuconostoc) and yeasts (Hancioglu and Karapinar, 1997; Zorba et al., 2003). Table 1 presents the microorganisms isolated from boza that have been reported by different studies. The beverage has a sour or sweet taste depending on the production method. It has a yellow color and a high viscosity. It has a unique acidic-alcohol odor, a low pH as well as a low alcohol content (Anonymous, TS 9778, 1992; Arici and Daglioglu,2002; Hancioglu and Karapinar, 1997;Zorba et al., 2003).Table 2 presents the composition of this beverage reported in the results of various studies.
L. monocytogenes is a gram-positive, non-spore forming, rod-shaped bacterium. The most incriminated food products include ready-to-eat food, meat, dairy, and aquaculture products (Meloni et al., 2009).Listeriosis can have a variety of symptoms including a mild fever, meningitis, septicemia, newborn infections and abortus (Salamina et al., 1996; Lokerse et al., 2016).It emerges with as high as 30% mortality rate in pregnant women, newborn infants, elderly persons, and individuals with suppressed immune system (McLauchlin, 1997).
Food products are formulated, produced and processed in order to minimize the growth of pathogenic microorganisms and target spoilage organisms. Nevertheless, the final product might contain pathogenic or spoilage organisms, or these organisms could grow in the formulation. They may also be already present in the product due to the contamination caused by the problems in the process or hygiene practices. For this reason, preventing the growth of undesirable organisms during the intended shelf life is necessary. There are few studies in the literature on the survival of pathogenic microorganisms in Boza. Also, there are no studies examining the survival of L. monocytogenes in the pH environment changing during the Boza fermentation. This study aims to determine the survival of L. monocytogenes during the production of boza and evaluate the microflora of the ripened boza as well as its inhibitory effect on the microflora on L. monocytogenes.
Table 1.Microorganisms isolated from boza as reported by studies
|
Hancioglu and Karapinar (2002)
|
LAB (Lactobacillus and Leuconostoc spp.), Yeast (Saccharomyces spp.) |
|
Botes et al. (2007) |
LAB (Lactobacillus spp.), Yeast (Candida spp. and Issatchenkia spp.) |
|
Gotcheva et al. (2000) |
LAB (Lactobacillus and Leuconostoc spp.) Yeast(Candida spp. and Geotrichum spp.) |
|
Kivanc et al. (2011) |
LAB (Lactococcus spp., Leuconostoc spp., Lactobacillus spp., Enterococcus spp., Pediococcus spp.) |
LAB: Lactic Acid Bacteria
Table 2.Composition of boza and the chemical values
|
Dry Matter (%) |
24.85 |
Zorba et al. (2003) |
|
Dry Matter (%) |
15.3–31.1 |
Yegin and Uren (2008) |
|
Protein, (%) |
1.06 |
Zorba et al. (2003) |
|
Protein, (%) |
0.50–0.99 |
Yegin and Uren (2008) |
|
Carbohydrates, (%) |
12.3 |
Zorba et al. (2003) |
|
pH |
3.16–4.02 |
Yegin and Uren (2008) |
|
Alcohol (ethanol), (%) |
0.13 |
Kose and Yucel (2003) |
Boza production
In this study, boza was produced under laboratory conditions in accordance with the traditional formulation. Firstly, a mixture of corn, wheat, and rice (at a ratio of 2:1:1, respectively), free from impurities and brans, was prepared. Sugar and starter culture was not added. This was continuously mixed for an hour with the addition of water (water/mixture ratio 1:1) until it became homogenous. Then, hot water (~ 65 °C) was added at 4:1 ratio. This new mixture was cooked and mixed for 45 minutes. Afterwards, it was cooled until reaching room temperature, diluted with water (2.5:1 ratio), and added sugar (20%), mixing it continuously (Hancioglu and Karapinar, 1997).
L. monocytogenes strains ATCC 13932 (4b strain) and ATCC 19111 (1/2a strain) at 70 log cfu/mL, (designated hour 0) were inoculated individually to the prepared beverage. Then, L. monocytogenes count was immediately calculated at zero hour. L. monocytogenes enumeration was conducted according to TS EN ISO 11290-2 (Anonymous, TS EN ISO 11290-2,2001.). This procedure was carried out at regular time intervals.
Microbiological Analyses
Boza sample (10 mL), was homogenized in 90 mL dilution medium that containing 0.1% peptone and 0.85% NaCl. For the isolation of Lactococcus, Enterococcus, and Lactobacillus strains, appropriate dilution rates were inoculated on M17 agar [Oxoid, added with 10% membrane filtered lactose solution (Oxoid, w/v)], Kanamycin esculin azide agar base (Oxoid, added with kanamycin supplement), and MRS agar (Oxoid), respectively. Diluted samples were incubated at 30ºC for 2 days under aerobic conditions on M17 agar, at 40ºC for 2 days under aerobic conditions on Kanamycin esculin agar, and at 37ºC for 2 days under anaerobic conditions on MRS agar. Three colonies from each medium were randomly taken, and purified in M17 broth [Oxoid, added with 10% membrane filtered lactose solution (Oxoid, w/v)], for Lactococci and Enterococci spp, and in MRS broth (Oxoid) for Lactobacilli (Tormoet al., 2015). This procedure was repeated twice. Gram staining and catalase tests were applied to the purified colonies, and Gram positive and catalase negative colonies were then stored in tubes containing 20% glycerol at -80ºC.
Experimental design
For the determination of phenotypic aspects, especially for differentiation of Enterococci and Lactococci, biochemical tests were performed. These tests were carried out at 10ºC, 15ºC, 40ºC and 45ºC in an optimal salt concentration of 2.5%, and at 30ºC in 2%, 4% and 6.5% salt concentrations using M17 and MRS broth media (Tormoet al., 2015). The pH of the boza was determined by using a pH meter (Inolab 7110, pH & oxygen meter).
Automated EcoRIribotyping
Based on the information above, the automated EcoRIribotyping identified 48 isolated suspected lactic acid bacteria. The standard method of the automated RiboPrinter® Microbial Characterization System was conducted for EcoRI ribotyping in accordance with the manufacturer’s instructions. To obtain the pure culture samples, MRS plates were incubated for 24–48 h at 30°C using appropriate colony samples. The microbial samples were analyzed following manufacturer’s instructions of the producer. The ribotype profiles used DuPont identification database DUP2003 to compare of the isolates with each other. When the corresponding pattern matched one of the patterns of the DuPont Identification Library with a similarity ≥ 0.85, that isolate was accepted as an identification. RiboPrinter™ (DuPont Qualicon) was used for dividing the isolates automatically into ribogroups based on the similarity of the respective ribotype patterns (Kivanc et al., 2011).
Determination of anti-Listerial activity
The isolated Enterococci strains were resuscitated at 30°C for 24 h in M17 broth. Further the strains were then grown at 30°C for 20 h under anaerobic conditions. For the determination of the anti listerial activity of isolates (whole culture broth), the study used an agar spot-on-a-lawn method (Todorov and Dicks, 2005) with the replacement of L. monocytogenes ATCC 19118 as the indicator strain. Wells of uniform diameter (6 mm) were cork bored in the agar and sealed with 15 mL of tempered soft agar. The cultures obtained from overnight cultures were applied to 4 different processes. (I): centrifuged at 2500 × g for 5 min. (II): neutralized by addition of 5 N NaOH to exclude the effects of organic acids. (III): treated with catalase (in 10 mM phosphate buffer, pH 7.0 to a final concentration of 1 mg/mL) for 30 min at 25 °C to exclude inhibitory effects of hydrogen peroxide and controls included phosphate buffer with and without catalase. (IV): added Proteinase K enzyme in 0.5 μg/mL final concentration and incubated for 4 h at 30 °C together with and without samples which including enzyme. Each of the solutions was sterilized with Millipore membrane filter (0.22 mm pore diameter) before loading to wells. Then, 20 µl samples were inoculated onto the surface of the BHI agar plates that had been inoculated with approximately 5 log cfu/ml of L. monocytogenes. The plates were incubated under aerobic conditions at 35°C for 15 h, and the diameters of clear zones of inhibition were calculated using the Computer Scanning System (Multi-ScanBase v. 14.02). The results were expressed in mm.PCR-identified Enterococcus faecium and Enterococcus faecalis strains obtained from the previous studies were used as positive control (Buyukyoruk et al., 2014).
After 48 hours of sharp decline in pH (from 6.55 to 3.52), the number of microorganisms also decreased significantly (from 5.60 log cfu/mL to 3.00 and/or 3.62 log cfu/mL) through ripening period. Although the pH value decreased to 3.70, inoculated L. monocytogenes strains were dominant in boza samples after 24 hours of storage at 4°C and reached up to 7.00 log cfu/ml. However, L. monocytogenes 1/2a and L. monocytogenes 4b strains could not be detected in microbiological tests after 72 hour Table 3 and Table 4.
Table 3. Hourly changes in Listeria monocytogenes*
|
Hours |
L. monocytogenes ATCC 13932ꭓ |
L. monocytogenes ATCC 19111ꭓ |
|
0 (zero) |
5.63 ± 0.035 |
5.61 ± 0.030 |
|
1 |
5.25 ± 0.050 |
5.39 ± 0.080 |
|
2 |
4.95 ± 0.040 |
5.67 ± 0.015 |
|
4 |
5.62 ± 0.030 |
6.34 ± 0.020 |
|
8 |
6.50 ± 0.015 |
7.3 ± 0.015 |
|
12 |
6.61 ± 0.020 |
7.2 ± 0.025 |
|
24 |
7.00 ± 0.020 |
6.57 ± 0.025 |
|
48 |
3.00 ± 0.020 |
3.62 ± 0.035 |
|
72 |
Not detected, <1 |
Not detected, <1 |
* : The average of five consecutive repetitions
ꭓ: logcfu/mL± standard variation
Table 4. Hourly changes in pH though of the ripened Boza*
|
Hours |
pH |
|
0 (zero) |
6.55 |
|
1 |
7.02 |
|
2 |
6.57 |
|
4 |
6.55 |
|
8 |
6.47 |
|
12 |
3.85 |
|
24 |
3.70 |
|
48 |
3.52 |
|
72 |
3.48 |
The ribotyping results revealed 23 Enterococcus faecium and 21 Enterococcus faecalis strains. (Fig 1 and Fig 2). Out of these isolates, 14 Enterococcus faecium and 8 Enterococcus faecalis strains had antilisterial activity. Enterococci showed an antilisterial activity in the form of a turbid zone round about the well and bacteriostatic activity against L. monocytogenes. Five of the strains were similar with Corynebacterium macginleyi (between 55-58% similarity), Xanthomonas campestris (58% similarity) and Corynebacterium simulans (54% similarity) from the Dupont Identification Library.
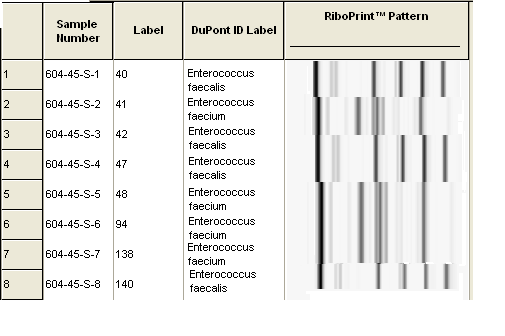
Figure 1. Dupont-ribotyping outputs of E. faecalis and E. faecium
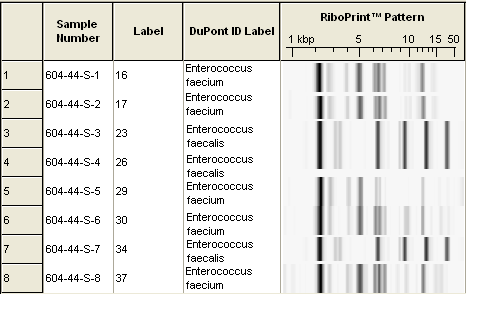
Figure 2. Dupont-ribotyping outputs of E. faecalis and E. faecium
Based on the literature research, no study was found regarding to L. monocytogenes in boza depending on pH changing and the food infection case due to boza consumption in Turkey.HoweverL. monocytogenes is an important pathogen due to some characteristics of bacteria. The organism is the causative agent of listeriosis, a severe disease with high hospitalization and case fatality rates. L.monocytogenes can survive and grow over a wide range of environmental conditions such as refrigeration temperatures, low pH and high salt concentration(Gandhi and Chikindas, 2007).Since this bacteria is one of the most dangerous microorganism indicated above, L. monocytogenes, should be considered as a remarkable source of public health problems, even its initial contamination level is low (Sarımehmetoğlu and Kaymaz,1994; Yıldırım and Sarımehmetoğlu, 2006; Şireli et al.,2008; Dikici et al., 2013; Zoz et al., 2017;Changcheng Liet al., 2017).
In this study, 1/2a and 4b serotypes were selected asreference L. monocytogenes strains, in which the previos one was foodborne and the latter was clinical strain. Although 13 serotypes have been described for L. monocytogenes (Shi et al.,2015), among these serovars, the 4b serotype is regarded to be responsible for the majority of human listeriosis cases, and serovar 1/2a is the most prevalent serovar in foods (Jamaliand Thong, 2014).
Boza harbours a diverse population of lactic acid bacteria. Diversity in bacterial and yeast populations from the different samples may be due to differences in raw materials, production processes and storage conditions. Examining the studies on the flora of boza (Table 1), only one study has reported the isolation of Enterococcus (Kivanc et al.,2011).The present study concluded that Enterococcus was the dominant flora in boza.
Hancioglu and Aktug-Gonul (1999), examined the survival and growth of E. coli O157:H7, Salmonella Typhimurium, and Staphylococcus aureus in boza, and found that E. coli O157:H7 was not inhibited and survived at pH 3.7 during a 32 hour fermentation. The growth of S. typhimurium and S. aureus were reported to be inhibited after 12 hour fermentation (pH <4.5). Guven K and Benlikaya N (2005), have reported that the acidic pH environment in boza had an important effect on the inhibition of Bacillus cereus.In another study, the antimicrobial effect of boza on various microorganisms was shown to be (in descending order) as total bacteria >E. coli>L. monocytogenes>S. pneumonia>B. cereus (Pehlivanoğlu et al.,2015).
Helland et al., (2004),reported that boza samples had a probioticeffect as they contained more than 106 cfu/mL viablemicroorganisms, which wassufficient for functional food in their study.Both lactic acid bacteria and yeasts were responsible for the spontaneous fermentation of many types of foods including boza (Arici and Daglioglu 2002). Cetinkaya and Soyutemiz (2004), have suggested that pathogens and spoilage microorganisms in fermented foods may be inhibited during the growth of lactic acid bacteria. In the present study, the inactivation of L. monocytogenes strains after a certain period of time was associated with the decrease in pH and the antilisterial activity of Enterococcus faecium and Enterococcus faecalis strains.
In this study, L. monocytogenesdid not surviveafter the production and ripening procedures of boza. However, good manufacturing practices should be considered in the production of ready-to-eat foods since this microorganism is abundant in nature and pottentially dangerous for public health.
Funding:This study was supported by Adnan Menderes University Scientific Research and Projects as Project No. VTF-11006.
Anonymous, 1992. TS 9778, Boza.
Anonymous, 2001. TS EN ISO 11290-2, Microbiology of food and animal feding stuffs- Horizontal method for the detection and enumeration of Listeriamonocytogenes- Part 2: Enumeration method.
Arici M, Daglioglu O, 2002. Boza: A lactic acid fermented cereal beverage as a traditional Turkish food. Food Rev Int 18: 39-48.
View ArticleBotes A, Todorov SD, von Mollendorff JW, Botha A, Dicks LMT, 2007. Identification of lactic acid bacteria and yeast from boza. Process Biochem 42: 267-270.
View ArticleBuyukyoruk, S, Naim DA, Emre YG, Devrim B, Pelin K, 2014. Species distribution, molecular characteristics and vancomycin resistance gene profiles of Enterococcus spp. isolates from home-made western cheeses in Turkey. Int J Dairy Technol 67 (1): 103-9.
View ArticleCetinkaya F, Soyutemiz GE, 2004. A study on survival of Listeria monocytogenes during manufacture and ripening of kashar cheese. Turk J Vet Anim Sci 28: 927-932.
Changcheng Li C, Huang L, Hwan CA, 2017. Effect of temperature and salt on thermal inactivation of Listeria monocytogenes in salmon roe. Food Control 73: 406-410.
View ArticleDikici A, Ilhak OI, Calicioglu M, 2013. Effects of essential oil compounds on survival of Listeria monocytogenes and Escherichia coli O157:H7 in çiğ köfte. Turk J Vet Anim Sci 37: 177-182.
Gandhi M., Chikindas ML, 2007. Listeria: A foodborne pathogen that knows how to survive. Int J Food Microbiol 113: 1-15. PMid:17010463
View Article PubMed/NCBIGotcheva V, Pandiella SS, Angelov A, Roshkova ZG, Weeb C, 2000. Microflora identification of the Bulgarian-cereal based fermented beverage Boza. Process Biochem36: 127-130. 00192-8
View ArticleGüven K, Benlikaya N, 2005. Acid pH produced by lactic acid bacteria prevent the growth of Bacillus cereus in boza, a traditional fermented Turkish beverage. J Food Safety 25: 98-108.
View ArticleHancioglu O, Karapinar M, 1997. Microflora of boza, a traditional fermented Turkish beverage. Int J Food Microbiol 35: 271-4. 01230-5
View ArticleHancioglu O, Aktug-Gonul S,1999. Biyoteknoloji Dergisi XI. Kukem-Biyoteknoloji Kongresi Ozel Sayisi, 23: 93-4.
Helland MH, Wicklund T, Narvhus JA, 2004. Growth and metabolism of selected strains of probiotic bacteria, in maize porridge with added malted barley. Int J Food Microbiol 91: 305-313. PMid:14984778
View Article PubMed/NCBIJamali H, Thong KL, 2014. Genotypic characterization and antimicrobial resistance of Listeria monocytogenes from ready-to-eat foods. Food Control44: 1-6.
View ArticleKivanc M, Yilmaz M,Cakır E, 2011. Isolation and identification of lactic acid bacteria from boza, and their microbial activity against several reporter strains. Turk J Biol 35: 313-324.
Kose E,Yucel U, 2003. Chemical composition of Boza. J Food Technol 1: 191-3.
Lokerse RFA, Maslowska-Corker KA, van de Wardt LC, Wijtzes T, 2016. Growth capacity of Listeria monocytogenes in ingredients of ready-to-eat salads. Food Control60: 338-345.
View ArticleMcLauchlin J, 1997. The pathogenicity of Listeria monocytogenes: a public health perspective. Rev Med Microbiol 8: 1-14.
View ArticleMeloni D, Galluzzo P, Mureddu A, Piras F, Griffiths M, Mazzette R, 2009. Listeria monocytogenes in RTE foods marketed in Italy: Prevalence and automated EcoRI ribotyping of the isolates. Int J Food Microbiol 129: 166-173. PMid:19100643
View Article PubMed/NCBISalamina G, Dalle Donne E, Niccolini A, Poda G, Cesaroni D, Bucci M, Fini R, Maldini M, Schuchat A, Swaminathan B, Bibb W, Rocourt J, Binkin N, Salmaso S, 1996. A foodborne outbreak of gastroenteritis involving Listeria monocytogenes. Epidemiol Infect 1996 117: 429-436. PMid:8972666 PMCid:PMC2271639
View Article PubMed/NCBISarımehmetoğlu B, Kaymaz Ş, 1994. Türk salamura beyaz peynirinde yapım ve olgunlaşma aşamalarının Listeria monocytogenes üzerine etkisi. Ankara Üniv Vet Fak Derg 41 (2): 234-242.
View ArticleShi W, Qingping W,Jumei Z,Moutong C, Zean Y, 2015. Prevalence, antibiotic resistance and genetic diversity of Listeria monocytogenes isolated from retail ready-to-eat foods in China. Food Control 47: 340-7.
View ArticleŞireli UT, Göncüoğlu M, Pehlivanlar S, 2008. Growth of Listeria monocytogenes in çiğ köfte (raw meat ball). Ankara Üniv Vet Fak Derg 55: 83-7.
Todorov SD, Dicks LMT, 2005. Lactobacillus plantarum isolated from molasses produces bacteriocins active against gram-negative bacteria. Enzyme Microb Tech 36: 318-326.
View ArticleTormo H, Lekhal DAH, Roques C, 2015. Phenotypic and genotypic characterization of lactic acid bacteria isolated from raw goat milk and effect of farming practices on the dominant species of lactic acid bacteria. Int J Food Microbiol 210: 9-15. PMid:26082325
View Article PubMed/NCBIYegin S, Uren A, 2008. Biogenic amine content of boza: A traditional cereal-based, fermented Turkish beverage. Food Chem 111: 983-7.
View ArticleYıldırım Y, Belgin Sarımehmetoğlu B, 2006. Beyaz peynir yapımında bazı probiyotik bakterilerin kullanılmasının Listeriamonocytogenes üzerine etkisi. Erciyes Üniv Vet Fak Derg 3(1): 1-7.
Zorba M, Hancioglu O, Genc M, Karapinar M, Ova G, 2003. The use of starter cultures in the fermentation of boza, a traditional Turkish beverage. Process Biochem 38: 1405-11. 00033-5
View ArticleZoz F, Grandvalet C, Lang E, Iaconelli C, Gervais P, Firmesse O, Guyot S, Beney L, 2017. Listeria monocytogenes ability to survive desiccation: Influence of serotype, origin, virulence, and genotype. Int J Food Microbiol 248: 82-9. PMid:28288399
View Article PubMed/NCBI